Week 9: Sea How Far We've Come: On Larval Development, Poster Presentations, and New Opportunities8/27/2017 Hello readers! Time has certainly flown here at OIMB, and it is hard to believe that this is the last week of my internship. Just as I've grown as a scientist here, so too my larvae have grown-- check out their progress in the slideshows below! Sea star larvae (Patiria miniata) Sea urchin larvae (Strongylocentrotus purpuratus) This week the interns had the opportunity to present our research findings in the form of a poster presentation to both members of the scientific community at OIMB, and also to visitors at the local Charleston Marine Life Center (CMLC). I found both presentation opportunities challenging and rewarding for different reasons. The CMLC opportunity pushed me to present my research findings in a way that was accessible (and interesting) to a non-scientific community, while during my presentation at OIMB I encountered research questions I was unable to fully answer (and I was reminded of all the exciting questions I've yet to answer in my research). While in many ways 9 weeks is a very short duration of time for research, I left both presentation opportunities aware of all that I was capable of learning and accomplishing in 9 weeks, and feeling immensely inspired by the research I'd seen my fellow interns accomplish during this time at OIMB.
While my time as an OIMB REU intern has come to a close, my work as a researcher is far from finished. I'm excited to share that this September I'll be participating in another NSF-funded marine biology research opportunity--this time studying marine ecology in Ireland! I will likely be blogging about my IRES experiences and will post a link when available. Until then, take care and thank you for reading my blog!
0 Comments
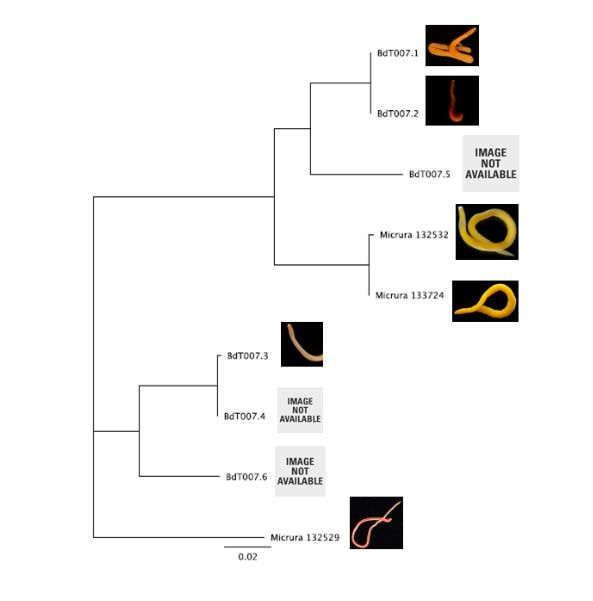
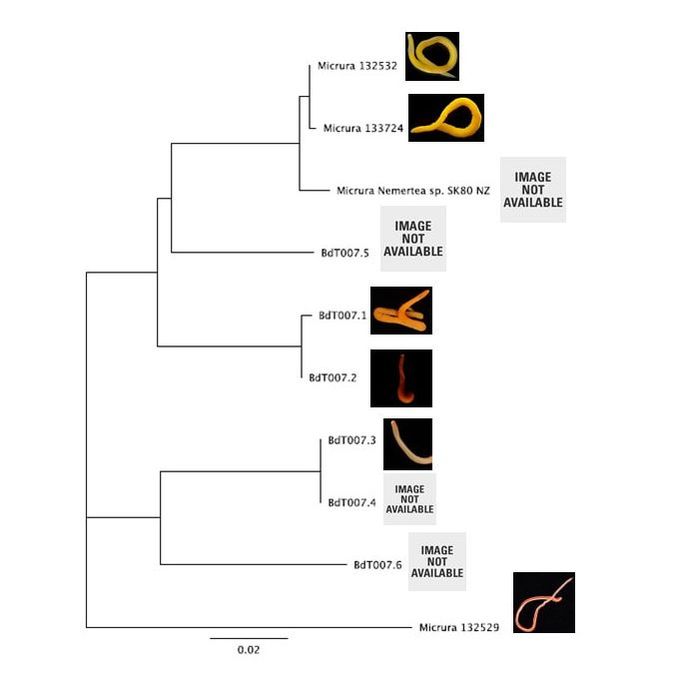
Hello readers! I am in my final week at OIMB (this is my second-to-last blog post), and while time certainly has flown by, I am amazed by all that has been accomplished during these past eight weeks. Despite all that I have learned during my internship, in many ways I feel that I am leaving OIMB with a greater certainty than before that I know very little about science; this is both a humbling and exciting realization, and it drives me to further pursue the fascinating subjects that I have been exposed to during my time here. As is often the case with research, nearly everything for my project came together very quickly at the last minute. On Thursday, I evaluated the quality of my newly-arrived sequences (most of which were high quality—yay!), trimmed the nucleotide ends of the sequences, and created consensus nucleotide sequences by pair-wise aligning the forward and reverse sequences for each gene. I then entered the gene nucleotide sequences into GenBank (an open access resource that matches sample sequences with over 150 million known nucleotide sequences) so as to contextualize my specimen sequences with established sequences. As expected, my sequences for the undescribed species did not fully match with any GenBank sequences; however, all of my sequences did most closely match with nucleotide sequences for the genes of fellow genus Micrura species, indicating that these specimens do indeed belong to genus Micrura. On Friday, Svetlana and I created phylogenetic trees for both the CO1 and 16S genes using a program called Geneious. After devoting weeks to re-extracting DNA, amplifying genes, troubleshooting amplification, then purifying and quantifying gene samples for sequencing, it was rewarding to finally see the fruit of my labor—these phylogenetic trees! As you can see from the distinct groupings (or clades), both trees provide evidence for the presence of five cryptic species, exceeding our preliminary expectations of three. Both trees group specimens BdT007.1 and BdT007.2 together as members of the same species, with BdT007.5 as a separate (but closely-related) species. Likewise, both trees group together BdT007.3 and BdT007.4 together as members of the same species, with BdT007.6 as a separate (but closely-related species). We also see that the closest specimens matched via GenBank to these species (Micrura 132432 and Micrura 133724) are closely-related to BdT007.1, BdT007.2 and BdT007.5 but in fact belong to a fifth separate species. We contextualized these relationships by adding an outgroup species (Micrura 132529) to the phylogenetic tree.
This findings illustrate the necessity of utilizing DNA barcoding when assessing species. Note how similar in appearance the yellow-hued Micrura 132432 and Micrura 133724 specimens are, but how orange-hued specimen BdT007.1 and red-hued BdT007.2 have noticeably different appearances from one another. Both pairs belong to the same respective species—this demonstrates that while morphological differences often serve as an indicator of the presence of multiple species, we see that this is not always the case. I am excited to share these findings with you, and am excited to share further findings as I delve more into the analysis of my research results. Comment below if you have questions, and look for larvae pictures next week! Hello readers! While I intended to devote this week’s blog post to describing the process of creating phylogenetic trees, it would feel disingenuous to describe this week’s exciting discoveries without acknowledging this week’s frustrations. Because my intention with this blog is to offer readers an opportunity to step into the world of research (with all its discoveries and failures), I'm devoting this week's blog post to sharing my struggles with frustration and insecurities as a scientist. I am learning here that while it is important to follow research deadlines as a scientist, science may not necessarily operate according to your deadlines, and research requires diligent planning but also patience, flexibility, and a sense of humor when you encounter failure. Early this week I received nucleotide sequences for the 16S and CO1 genes I had amplified, purified, and quantified last week, and was disappointed to discover that many of the sequences were too low of quality to be utilized in creating phylogenetic trees. While this discovery was by no means disastrous (I have enough extracted DNA to re-amplify, purify, and quantify the genes from the specimen), this discovery did set my research back nearly a week behind schedule. I had been eagerly anticipating analyzing the phylogenetic relationships between the nemertean specimen (are they all members of the same species or are some of them separate, cryptic species?), and I found myself internalizing my frustrations with this setback through self-blame (were the low-quality sequences a result of simple errors I’d made during amplification or purification?). While it is important to critically analyze possible sources of error, it is also important to approach troubleshooting in a productive manner, and upon reflection I realized that my ruminations over my procedures were causing me to feel less engaged in my project and less confident in my abilities as a scientist. I realized I was so fixated on controlling a small component of my project that I was diminishing my curiosity as a researcher. I believe that curiosity (followed by lots of hard work) drives discovery, enables researchers to make crucial conceptual connections, and makes research a joyful (and worthwhile) experience, and I realized I needed to revive my curiosity for my project. I did so in a rather unexpected way—by attending a public lecture on the social behavior of orcas. While I do admittedly have a soft spot for charismatic megafauna, while listening to the lecture I found myself most fascinated with the species delimitation aspects of the research. The researcher described different “ecotypes” of the same orca species he’d observed—all of whom were reproductively isolated from one another and differed in their morphology, prey choices, and social behaviors—and I found my mind reeling with questions about how a species is defined. Can differing social behaviors between reproductively-isolated pods indicate the presence of multiple species? Were different ecotype members physically incapable of successfully producing offspring together, or merely socially-disinterested? I found myself longing to answer these questions by uncovering the evolutionary histories of these ecotypes via genetics, and I wondered what their phylogenetic trees would look like. My curiosity about species delimitation reminded me of my motivation for researching putative cryptic species—and I returned to my project with regained enthusiasm. My experiences this week taught me that while setbacks are an unavoidable aspect of research, setbacks also provide an opportunity to reflect upon one’s mindset as a researcher, and I feel incredibly fortunate to have the opportunity to learn and grow as a researcher in an academic environment that is so supportive and intellectually-stimulating. My nemertean phylogenetic trees are (gradually) coming together, and I am excited to share my findings with you next week. Until then, check out how much my sea urchin (S. purpuratus) larvae have developed, and comment below with questions! |
Author“Instructions for living a life./ Pay attention./ Be astonished./ Tell about it.” (Mary Oliver) Archives |

 RSS Feed
RSS Feed