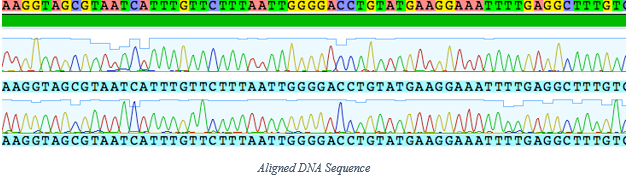
Week 8 To finish up my discussion regarding the processes involved in my research, this week I will cover sequencing the DNA. Once the DNA has been purified and quantified (week 7 blog post), we send our samples to a company called Sequetech located in Mountain View, CA in order to be sequenced. Once these sequences are ready, we need to analyze them and clean them up a bit. To do this, we use Geneious®, a sequencing analysis software. We first “trim off” the primer section of the DNA, then align the two complimentary strands to look for any discrepancies. Once the two complimentary strands are cleaned up and properly aligned, we can search for any matches to that particular sequence using GenBank®, a collection of publicly available DNA sequences. This data allows us to build phylogenetic trees and observe genetic relationships between species that we are studying. From here we are able to see if we have a species match, an undescribed species, or possibly even cryptic species. The past few blog entries have summed up the majority of my specific research, but the data that has been collected will be further analyzed and used as part of a much larger body of research long after I am finished with this program. I am excited to have the ability to follow the progress of this research into the future. 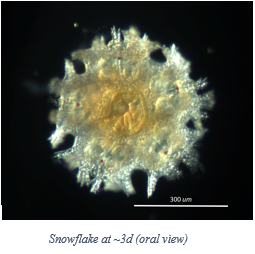 In week 2 I had mentioned that we had started some cultures of various organisms as a kind of side project, and this last week we have been able to see the results as our larvae have begun metamorphosing into their juvenile forms. The one that I am most proud of though, is our Patiria miniata larvae, as this is one of the more difficult organisms to induce to settle in a lab setting. When this program first began, I was informed that this had not yet been done in our lab, which of course I took as a personal challenge. I am excited to say that we have been wildly successful in getting our Patiria to settle, and I would like to introduce to you Snowflake: the first of his name, king of the larval culture, ruler of the Pyrex dish, and possessor of many tube feet.
0 Comments
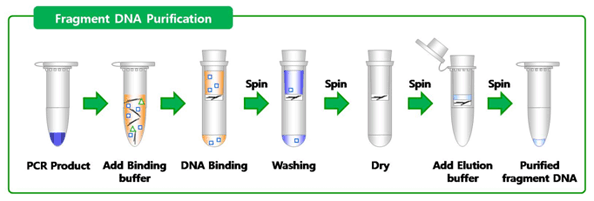
Week 7 This week I will cover the basics of DNA purification and quantification. Once we have amplified the target DNA using PCR, we need to purify the product (by removing primers, unincorporated nucleotides, polymerase, and salts) and quantify it in order to get a general idea of how much DNA is in the sample; if there is not enough DNA or too much DNA sequencing may fail. I used a commercial kit (Promega Wizard® SV Gel and PCR Clean-Up System) to purify our samples. First, a membrane binding solution is mixed with the PCR product and the mixture is pipetted into a spin column containing a silica membrane placed inside a collection tube; this assembly is then centrifuged in order to bind the DNA to the membrane (while everything else flows through into the collection tube). An ethanol-based wash solution is then added and pulled through the membrane by centrifuging in order to clean the filter. Once the sample is bound to the membrane and washed of contaminants, the spin column is placed into a fresh collection tube, and DNA is eluted by adding nuclease-free H20 to the filter, and centrifuging the assembly. This purified sample is then quantified using gel electrophoresis that I mentioned previously (see my previous blog post week-6.html). With this round of gel electrophoresis however, the sample is compared to a standard called Low Mass Ladder (which contains a mixture of fragments of DNA of known size and quantity) in order to estimate the quantity of DNA contained within the sample. Once the sample is purified and quantified, it is ready to send off for sequencing. The process of sequencing will be the focus of next week’s blog entry. (Photo courtesy of Bioneer Pacific) With only two weeks left in the program, we have officially run out of time for our usual weekly extracurricular activities. Focus has now shifted to compiling the data and preparing to present it to various audiences. We are all ready to see our hard work turn into something tangible, and the excitement around campus is palpable (punctuated with healthy doses of anxiety).
Week 6 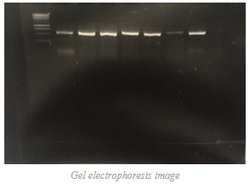 Week 6 To continue on with the breakdown of the processes involved in my research, I will explain the next process involved: gel electrophoresis. After completing PCR, the (hopefully) amplified DNA is known as PCR product. This PCR product is then run on a gel in order to verify that the amplification process was indeed successful, and to confirm that the amplified product is of the expected length. To do this, we use a 1% agarose gel with 0.01% ethidium bromide. Agarose is heated to liquefy it and then poured into a mold to form a 3D matrix with wells for the samples. The ethidium bromide fluoresces under UV light when bound to the DNA. Once the gel is cooled and set, a “ladder” is deposited into one of the wells to use as a reference. A ladder is a mixture of PCR products of known size. By comparing the position of our samples on the gel to the position of various bands in the ladder we can estimate the size of our PCR products (thus determining whether they likely represent the gene we were targeting or not). The gel is positioned inside of a gel rig containing a buffer solution. The migration tank has a negative charged electrode (cathode) and a positively charged electrode (anode) at opposite ends. The phosphate backbone of DNA is negatively charged, and so travels toward the anode in the presence of electrical current. Longer DNA fragments travel through the gel slower than the shorter fragments, so we can gauge the length of the DNA fragments (PCR products) based on how far they travel in the gel compared to the control (ladder). Once we have used this process to verify that PCR amplification was successful (a band of expected size and sufficient brightness is present), we can move on to PCR purification which I will explain in more detail next week.  As I have mentioned before, each week we try to get away and do something fun as a group, and this week was no exception. We spent this past Sunday frolicking in one of my favorite spots in Oregon, Coquille River Falls. This is one of the only spots that I am aware of in the area where the falls are not only accessible, but the pools are beautiful and deep enough to swim in. The weather was sunny and warm, and the water was a gorgeous, crystal clear deep green. We also had a rare opportunity to view a deceased juvenile gray whale that had washed up on a local beach. This may not be everyone’s cup of tea, but it was a great opportunity for students who are studying marine biology. Week 5 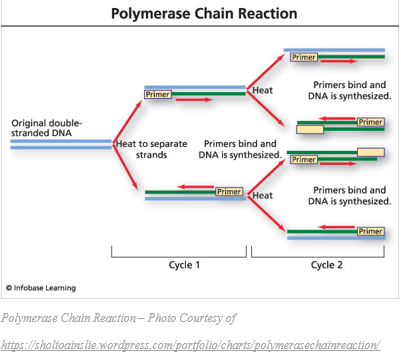 Last week I explained why I am doing the research that I am doing, and the significance of such research. In week 3 I promised to provide some details on the procedures involved in this research, so this week I will explain a little about polymerase chain reaction (PCR). PCR is the process used to amplify fragments of DNA that have been extracted from a specimen in order to make millions of copies to prepare for sequencing. To do this, we choose a particular gene or fragment of a gene to target. In this case, it is the gene that encodes a large subunit ribosomal RNA, 16S. This gene region is adjacent to highly conserved regions of DNA, and therefore we can use “universal” primers to start the process of transcription (Palumbi et al., 1991). Besides primers, we also need the nucleotide building blocks of DNA, DNA polymerase that can withstand high temperatures, and of course, a template strand of DNA that we want to copy. Once these are all mixed in the correct quantities, we can begin the process of PCR. Amplification happens through repeated cycles of heating and cooling; heating to 95ºC in order to denature or separate the two strands of DNA in the template, followed by cooling to 50ºC to allow the primers to anneal to the separated strands of DNA, then warming to 72ºC to allow the DNA polymerase to elongate the DNA. This process is repeated between 25 and 40 times, producing up to a million copies of the DNA (Rapley, 1998). This week officially marks the halfway point of the OIMB REU program for 2017; where has the time gone? With each day that passes, I am increasingly thankful to have this opportunity to gain valuable research experience that will greatly benefit me as I move forward into a science-based career. This program has given me the confidence that I needed to finalize my degree path, and because of this program, I have decided to pursue a minor in biology in addition to my bachelor’s degree in natural resources. I am looking forward to what the second half will bring. This past Friday, the REU interns were treated to a tour of the biology department at the main campus of UO in Eugene, where we were allowed to observe multiple studies across a variety of labs. While on campus, we also spent time exploring the UO Museum of Natural and Cultural History where we observed features of geological processes across the state of Oregon, artifacts used by early people in the area, as well as some thought-provoking photographic exhibits; I would recommend this stop to anyone going to the Eugene area. Palumbi S, Martin A, Romano S, McMillan WO, Stice L, Grabowski G. (1991) The simple fools guide to PCR Version 2.0. Honolulu, HI: Department of Zoology and Kewalo Marine Laboratory, University of Hawaii. Rapley, R., & Walker, John M. (1998). Molecular biomethods handbook. Totowa, N.J.: Humana Press. WEek 4  For this week's blog post, I have decided to give a bit more detail into the research that I will be doing over the next few weeks. To do this, I will provide snippets of my recent research proposal to give you some insight into why I will be doing this specific research. Introduction
I propose to assess the species diversity of phylum Nemertea (ribbon worms) of Bocas del Toro, Panama. Estimates vary, but it is clear that most species on Earth remain undescribed (Appeltans et al., 2012; Mora et al., 2011). Some taxa and regions in the world are better studied than others, and nemerteans are one of the lesser known phyla of marine invertebrates. In particular, biodiversity of this group in the tropical regions is poorly known. There are only 16 species of nemerteans reported from Bocas del Toro, most of which are undescribed (Collin et al., 2005). Recent estimates suggest that the actual diversity is at least five times that number (Maslakova, pers. comm.). Classical species identification relies on morphological characteristics that may not be a reliable means of distinguishing between cryptic species or newly diverged species. For instance, a previously known species of nemertean, Micrura alaskensis, has recently been found to be five separate cryptic species (Hiebert & Maslakova, 2015). DNA barcoding can be used to identify species on a molecular level when morphological characteristics are inadequate. Significance This research will help to document and increase biodiversity of a phylum that before now has been severely understudied. By combining DNA barcoding with classical morphological species identification, we can identify species at a much faster rate than ever before. This is especially important with increasing extinction rates due to anthropogenic disturbances, with extinction rates becoming higher than discovery rates of new species (Miloslavich et al., 2010). DNA barcoding also allows for matching of larval forms to adult forms which is not easily done using morphological characteristics. Identifying larval forms helps understanding of life phases, as well as increases biodiversity and understanding by finding larval forms when adults are unavailable. My research will continue over the next 5 weeks until Friday, August 11th, with approximately 45 samples completed each week using the previously mentioned processes. Presentation of my results will be on Monday, August 14th at Charleston Marine Life Center in Charleston, Oregon. The research will be ongoing in the lab after I am finished with the OIMB REU program. Week 3  Per my promise in last week’s blog, I will start this week by giving you some highlights of our dredging trip on the Pluteus, OIMB’s research vessel. On Saturday, all of the REU interns along with our fearless leaders, Maya, Richard, and MacKenna, boarded the Pluteus at 8:00am to embark on a journey out to sea (a first for some of the students) to see what amazing organisms we might be able to find by dredging. Dredging is the process of collecting benthic organisms by dragging a basket along the sea floor for a short distance. Our first dredge proved the most fruitful, we pulled up a plethora of fascinating creatures including sea cucumbers (Cucumaria miniata), some sea stars that appeared to be from another planet known as basket stars (Gorgonocephalus eucnemis), and even a small octopus (which wouldn’t hold still long enough for me to get a good photo). On a second dredge over sandy substratum, we brought up thousands of juvenile sand dollars (Dendraster excentricus), which at their size would be more easily pictured if thought of as “sand dimes.” A few unlucky students spent part of the voyage staring at the horizon, attempting to hold down their breakfast, a malady that I was not completely immune to myself, despite having undertaken many such trips in the past.  After receiving in-depth and hands-on training during my first two weeks, this week I was given free reign, starting Monday, to begin working in earnest on “The Panama Project”. My work on this project will center around DNA barcoding the nemertean ribbon worms collected from Bocas del Toro, Panama. I will be giving more in depth descriptions of these procedures in the following weeks, as I can’t give away all the amazing stuff in just one blog post. As with last week, we interspersed our hard work with some play including a fourth of July picnic at Sunset Bay state park and more trips to collect food for our specimens from the docks. week 2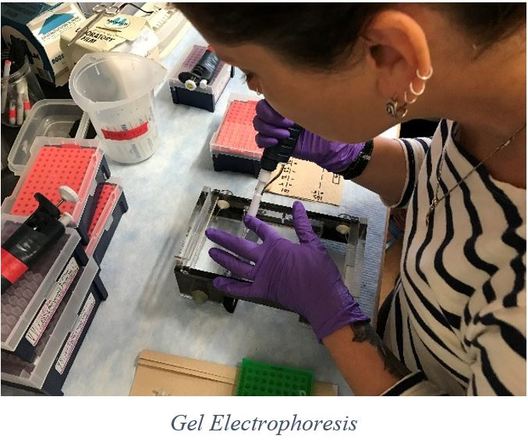 These past two weeks have been a whirlwind of training, hands-on research, and team building. After becoming familiar with the lab in which I will be doing my research, I was able to gain some experience with the myriad of equipment that I will be using through hands-on training. The bulk of my research will be dedicated to DNA barcoding of Nemertean ribbon worms (a phylum of marine worms in which many species are undescribed). While I may to refer to this as “my research,” I am working closely with a dedicated team of graduate students that are attending at OIMB, as well as my mentor, Svetlana Maslakova. The support and training that they have provided so far is invaluable. To become familiar with the processes involved, I was provided with specimens, both live and preserved, in which to work with. The first process involved extracting DNA from the specimens, after which the DNA had to be amplified using polymerase chain reaction. Following amplification of the DNA, we then verified our samples using gel electrophoresis. Once the samples were verified, we then purified and quantified the samples in order to ensure there were adequate quantities to send off for sequencing.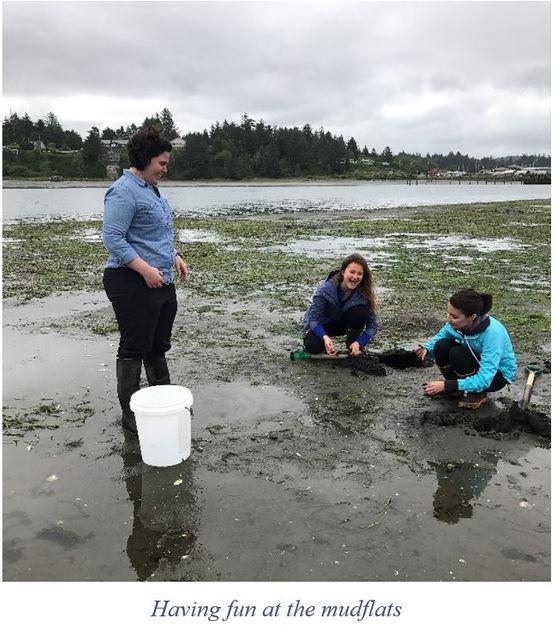  While this all may sound very technical, my time in the program so far has not been all work and no play. Along with the research that we are doing in the lab, we have also spent time out in the field collecting specimens and learning about the broad spectrum of organisms that reside in the local intertidal areas. In the lab, we have also started cultures of some of these in order to observe fertilization and larval stages. Some of the species that we have cultured include Pateria miniata (bat stars), Strongylocentrotus purpuratus (purple sea urchins), Phoronopsis harmeri (horseshoe worms), and Maculaura alaskensis (ribbon worms). As a team, we have also gone on camping trips locally, enjoyed some downtime with a picnic at the beach, and have a dredging trip planned for this weekend on OIMB’s research vessel, the Pluteus. Look for details of this trip in next week’s blog! |
AuthorMy name is Becky, and I have been local to the Coos Bay and Charleston area for two years with my boyfriend David, and my dog Mojo. I moved to this area to complete my associate's degree at Southwestern Oregon Community College in anticipation of transferring to a four-year university. I now attend online at Oregon State University and I am planning a move to Bend, OR in August to continue at the OSU Cascades campus in order to finish my bachelor's degree in either natural resources or biology. Archives |

 RSS Feed
RSS Feed