Week 9: Sea How Far We've Come: On Larval Development, Poster Presentations, and New Opportunities8/27/2017 Hello readers! Time has certainly flown here at OIMB, and it is hard to believe that this is the last week of my internship. Just as I've grown as a scientist here, so too my larvae have grown-- check out their progress in the slideshows below! Sea star larvae (Patiria miniata) Sea urchin larvae (Strongylocentrotus purpuratus) This week the interns had the opportunity to present our research findings in the form of a poster presentation to both members of the scientific community at OIMB, and also to visitors at the local Charleston Marine Life Center (CMLC). I found both presentation opportunities challenging and rewarding for different reasons. The CMLC opportunity pushed me to present my research findings in a way that was accessible (and interesting) to a non-scientific community, while during my presentation at OIMB I encountered research questions I was unable to fully answer (and I was reminded of all the exciting questions I've yet to answer in my research). While in many ways 9 weeks is a very short duration of time for research, I left both presentation opportunities aware of all that I was capable of learning and accomplishing in 9 weeks, and feeling immensely inspired by the research I'd seen my fellow interns accomplish during this time at OIMB.
While my time as an OIMB REU intern has come to a close, my work as a researcher is far from finished. I'm excited to share that this September I'll be participating in another NSF-funded marine biology research opportunity--this time studying marine ecology in Ireland! I will likely be blogging about my IRES experiences and will post a link when available. Until then, take care and thank you for reading my blog!
0 Comments
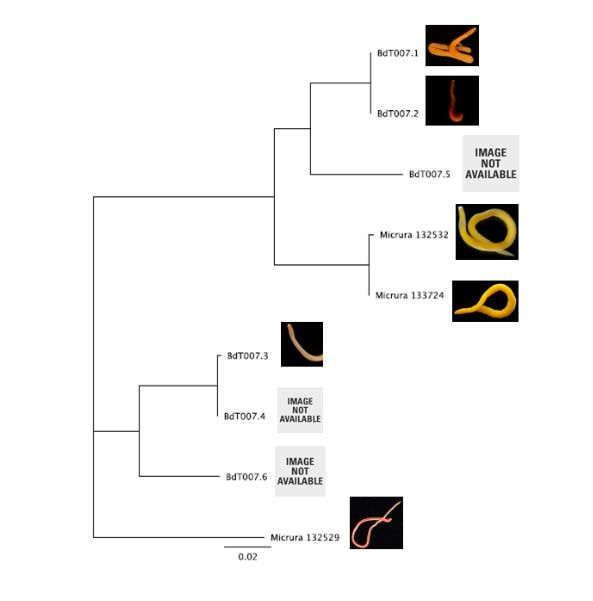
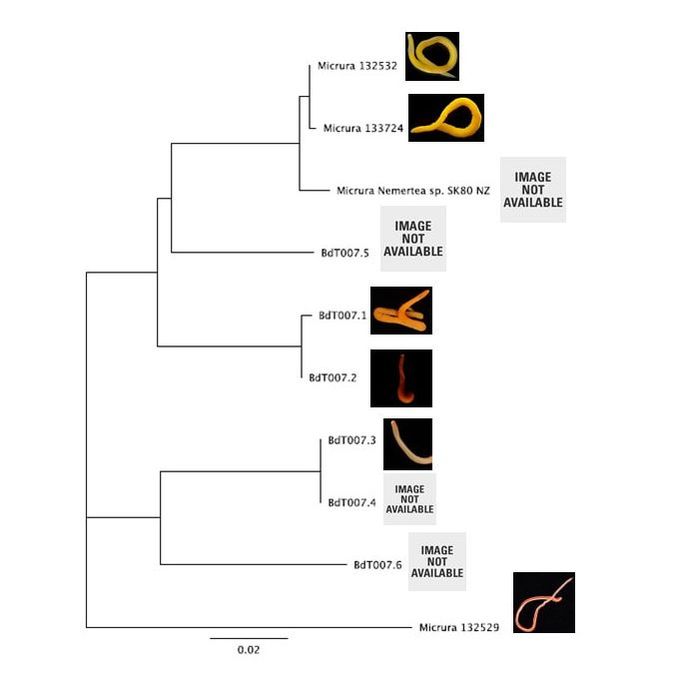
Hello readers! I am in my final week at OIMB (this is my second-to-last blog post), and while time certainly has flown by, I am amazed by all that has been accomplished during these past eight weeks. Despite all that I have learned during my internship, in many ways I feel that I am leaving OIMB with a greater certainty than before that I know very little about science; this is both a humbling and exciting realization, and it drives me to further pursue the fascinating subjects that I have been exposed to during my time here. As is often the case with research, nearly everything for my project came together very quickly at the last minute. On Thursday, I evaluated the quality of my newly-arrived sequences (most of which were high quality—yay!), trimmed the nucleotide ends of the sequences, and created consensus nucleotide sequences by pair-wise aligning the forward and reverse sequences for each gene. I then entered the gene nucleotide sequences into GenBank (an open access resource that matches sample sequences with over 150 million known nucleotide sequences) so as to contextualize my specimen sequences with established sequences. As expected, my sequences for the undescribed species did not fully match with any GenBank sequences; however, all of my sequences did most closely match with nucleotide sequences for the genes of fellow genus Micrura species, indicating that these specimens do indeed belong to genus Micrura. On Friday, Svetlana and I created phylogenetic trees for both the CO1 and 16S genes using a program called Geneious. After devoting weeks to re-extracting DNA, amplifying genes, troubleshooting amplification, then purifying and quantifying gene samples for sequencing, it was rewarding to finally see the fruit of my labor—these phylogenetic trees! As you can see from the distinct groupings (or clades), both trees provide evidence for the presence of five cryptic species, exceeding our preliminary expectations of three. Both trees group specimens BdT007.1 and BdT007.2 together as members of the same species, with BdT007.5 as a separate (but closely-related) species. Likewise, both trees group together BdT007.3 and BdT007.4 together as members of the same species, with BdT007.6 as a separate (but closely-related species). We also see that the closest specimens matched via GenBank to these species (Micrura 132432 and Micrura 133724) are closely-related to BdT007.1, BdT007.2 and BdT007.5 but in fact belong to a fifth separate species. We contextualized these relationships by adding an outgroup species (Micrura 132529) to the phylogenetic tree.
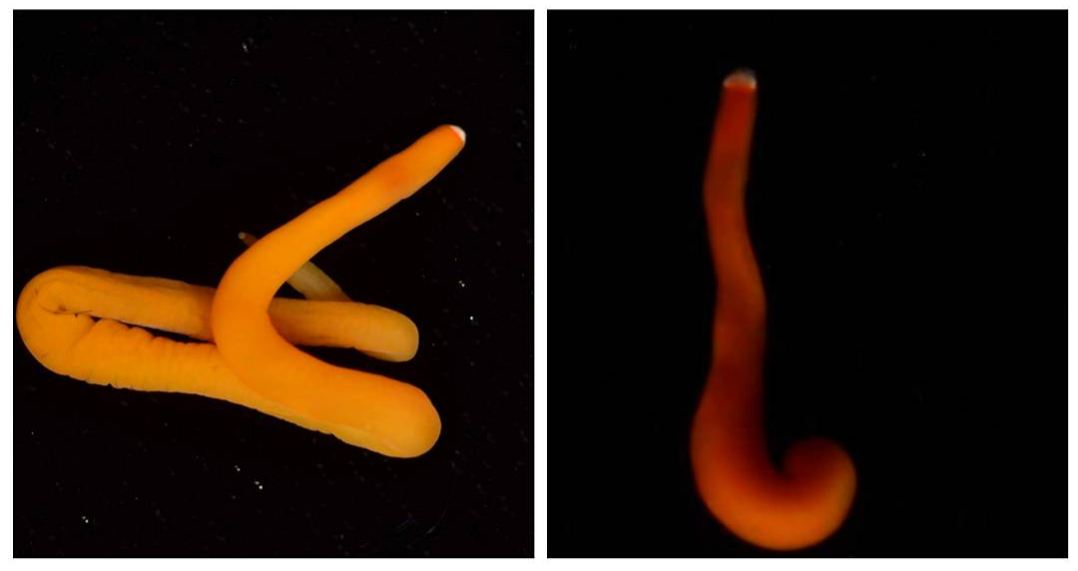
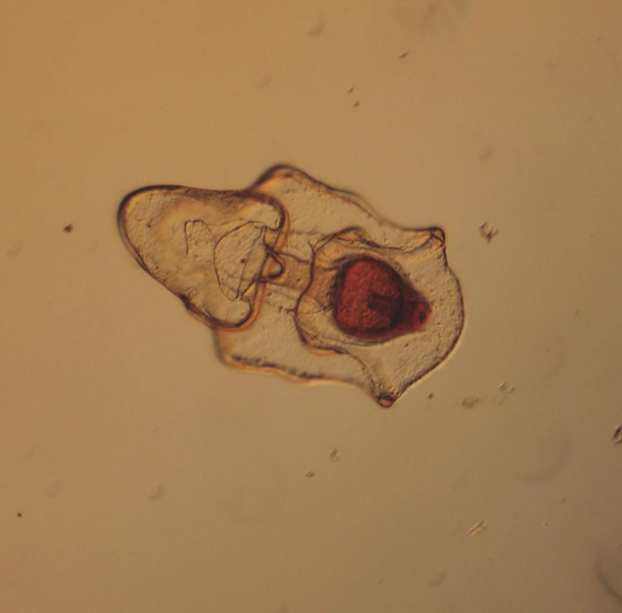
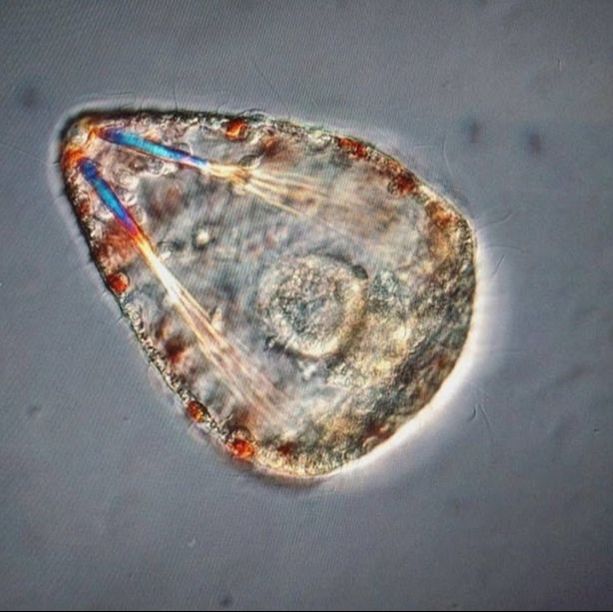
This findings illustrate the necessity of utilizing DNA barcoding when assessing species. Note how similar in appearance the yellow-hued Micrura 132432 and Micrura 133724 specimens are, but how orange-hued specimen BdT007.1 and red-hued BdT007.2 have noticeably different appearances from one another. Both pairs belong to the same respective species—this demonstrates that while morphological differences often serve as an indicator of the presence of multiple species, we see that this is not always the case. I am excited to share these findings with you, and am excited to share further findings as I delve more into the analysis of my research results. Comment below if you have questions, and look for larvae pictures next week! Hello readers! While I intended to devote this week’s blog post to describing the process of creating phylogenetic trees, it would feel disingenuous to describe this week’s exciting discoveries without acknowledging this week’s frustrations. Because my intention with this blog is to offer readers an opportunity to step into the world of research (with all its discoveries and failures), I'm devoting this week's blog post to sharing my struggles with frustration and insecurities as a scientist. I am learning here that while it is important to follow research deadlines as a scientist, science may not necessarily operate according to your deadlines, and research requires diligent planning but also patience, flexibility, and a sense of humor when you encounter failure. Early this week I received nucleotide sequences for the 16S and CO1 genes I had amplified, purified, and quantified last week, and was disappointed to discover that many of the sequences were too low of quality to be utilized in creating phylogenetic trees. While this discovery was by no means disastrous (I have enough extracted DNA to re-amplify, purify, and quantify the genes from the specimen), this discovery did set my research back nearly a week behind schedule. I had been eagerly anticipating analyzing the phylogenetic relationships between the nemertean specimen (are they all members of the same species or are some of them separate, cryptic species?), and I found myself internalizing my frustrations with this setback through self-blame (were the low-quality sequences a result of simple errors I’d made during amplification or purification?). While it is important to critically analyze possible sources of error, it is also important to approach troubleshooting in a productive manner, and upon reflection I realized that my ruminations over my procedures were causing me to feel less engaged in my project and less confident in my abilities as a scientist. I realized I was so fixated on controlling a small component of my project that I was diminishing my curiosity as a researcher. I believe that curiosity (followed by lots of hard work) drives discovery, enables researchers to make crucial conceptual connections, and makes research a joyful (and worthwhile) experience, and I realized I needed to revive my curiosity for my project. I did so in a rather unexpected way—by attending a public lecture on the social behavior of orcas. While I do admittedly have a soft spot for charismatic megafauna, while listening to the lecture I found myself most fascinated with the species delimitation aspects of the research. The researcher described different “ecotypes” of the same orca species he’d observed—all of whom were reproductively isolated from one another and differed in their morphology, prey choices, and social behaviors—and I found my mind reeling with questions about how a species is defined. Can differing social behaviors between reproductively-isolated pods indicate the presence of multiple species? Were different ecotype members physically incapable of successfully producing offspring together, or merely socially-disinterested? I found myself longing to answer these questions by uncovering the evolutionary histories of these ecotypes via genetics, and I wondered what their phylogenetic trees would look like. My curiosity about species delimitation reminded me of my motivation for researching putative cryptic species—and I returned to my project with regained enthusiasm. My experiences this week taught me that while setbacks are an unavoidable aspect of research, setbacks also provide an opportunity to reflect upon one’s mindset as a researcher, and I feel incredibly fortunate to have the opportunity to learn and grow as a researcher in an academic environment that is so supportive and intellectually-stimulating. My nemertean phylogenetic trees are (gradually) coming together, and I am excited to share my findings with you next week. Until then, check out how much my sea urchin (S. purpuratus) larvae have developed, and comment below with questions! Hello readers! Given that I spent much of this week waiting for the arrival of my sequence results from my putative cryptic species samples, I’m devoting this week’s blog post to describing my experiences collecting nemerteans in the local mudflats. Engaging in hands-on fieldwork is one of the primary reasons I love to study science, and after weeks of studying nemertean DNA in the lab, I found it highly satisfying to observe live nemerteans in their natural habitat. We arrived at the mudflats midmorning with the primary objective of collecting a specific nemertean species (M. alaskensis) for a graduate student’s research on egg parasitism—however, this objective didn’t stop us from having fun looking at all the worms (nemertean and non-nemertean) we found in the mudflats! I was surprised by the number of graduate students who volunteered for this excursion, and I was reminded of how distinct the OIMB community is in its enthusiastic devotion to research. In my experience, it’s not often you find a large number of people who’d gladly spend their morning digging in mud for worms, and I feel immensely lucky to be part of a community of scientists who (while engaged in demanding projects of their own) consistently find the time, energy and curiosity to engage in data collection for research done by others. We located nemerteans in the mudflats by digging into the sediment with shovels. Despite their somewhat desolate appearance, mudflats are diverse ecosystems. Migratory shorebirds hopped about, hungrily eyeing the sediment surface for indications of burrowing clams, their efforts perhaps less successful than those of the local residents armed with shovels and buckets. We students, however, had no competition for the mudflat’s nemerteans (which have few natural predators due to their toxicity). Our biggest challenge was correctly identifying the wriggling, mud-covered worms, and extracting them from the sediment without damaging their fragile bodies. An added excitement was observing how nemerteans, when threatened, extend their proboscis. Check out the length of the (white) proboscis on this nemertean! We transported our collected nemerteans back to the lab in small test tubes, gently rinsed the sediment off their bodies, then placed them in a container of filtered seawater. I am fascinated by nemertean locomotion—look at how gracefully this Cerebratulus cf. marginatus glides through the water! This excursion incentivized me to study nemertean taxonomy. In a classroom setting I often struggle with motivating myself to memorize taxonomy; in the field however, I found myself wishing that I could identify and describe all the nemerteans I was observing (luckily, I was with knowledgeable graduate students). While I’m still struggling to train my eyes to recognize identifying characteristics in nemertean morphology, this week I’m challenging myself to study nemertean species local to the Pacific Northwest in preparation for future mudflat excursions. Next week I’ll describe the process of analyzing gene sequences and update you with pictures of my larvae (they’ve gotten so big!). Until then, take care, and comment below if you have questions. Hello readers! As promised, I’m devoting this week’s blog post to describing my primary research project, identifying and phylogenetically categorizing cryptic ribbon worms from Panama. It’s been an incredible experience to study ribbon worms with renowned expert, Svetlana Maslakova, and I hope that this week’s blog post (and included videos) will inspire you to join me in my new-found fascination with these “pythons of the sea!” So… what are ribbon worms? Ribbon worms (nemerteans) are a highly-diverse phylum of marine-dwelling, unsegmented worms characterized by their proboscis, an oral appendage used to capture and stun prey (check out this video of a ribbon worm with a branching probisis: https://youtu.be/_keb7YpmAls). Ribbon worms range in length from 2 millimeters (genus Carcinomertes) to over a 100 feet (Lineus longissimus), most have planktonic larval stages and some undergo “catastrophic metamorphosis,” where the juvenile emerges from and subsequently consumes its larval body (and you thought puberty was rough?!). Check out Svetlana’s video of this incredible process: https://youtu.be/0Kt9BRcNB5w. Recent estimates suggest that the majority of species on Earth remain undescribed. The fraction of undescribed species is largest in understudied taxa, such as ribbon worms. Some 1300 species are described, but the true diversity is likely several times that number. Studies of ribbon worms are further complicated by the presence of cryptic species, which are are morphologically-indistinguishable but identifiable as separate species via DNA-barcoding (comparisons of select genes). Cryptic species pose a challenge to conservation efforts because while visually similar, cryptic species may have different ecological roles and thus require different management strategies. Cryptic species also pose a challenge to biodiversity studies, because a population count for an endangered species may in fact describe multiple cryptic species (meaning there are more endangered species, with lower population numbers). DNA barcoding has been indispensable for the discovery and characterization of cryptic species, and I will be utilizing such methods for my research. The putative cryptic species complex I’m researching was discovered by Svetlana and her graduate student Nicole Moss during a 2016 expedition to Bocas del Toro, Panama. On site, ribbon worm specimens were observed, photographed, and had tissue removed for DNA extraction (check out Svetlana’s video on how ribbon worms are collected from their environment: https://youtu.be/CX-C-vdEK0Y); during this process, the researchers observed subtle color variation between individuals of what looked like a single species, indicating the potential for a cryptic species complex (see pictures below). Preliminary DNA-sequencing efforts confirmed that this species may comprise several cryptic closely related species. My research seeks to determine whether or not these individuals belong to separate species (and how many), and my work begins with the extracted DNA. Currently, I am PCR-amplifying fragments of two mitochondrial genes: Cytochrome Oxidase Subunit I and 16S rDNA from the DNA samples extracted from six individuals. Next week I plan to send my purified PCR products to a sequencing company. Once I’ve received the nucleotide sequences, I will build phylogenetic trees and calculate sequence divergence between individuals. I feel as though I am methodically solving a mystery through my research here, and it is truly thrilling! Equally thrilling is the knowledge that my research has the potential to serve as a resource for future conservation or biodiversity studies in Bocas del Toro, Panama. There are more fascinating aspects of ribbon worms than I can possibly cover in one blog post, but I hope I’ve inspired you to learn more about this underappreciated phylum. Please comment below if you have questions, and check out Svetlana’s website for further information http://pages.uoregon.edu/svetlana/iWeb/Home/Nemerteans.html. Hello readers! Given the exciting morphological progressions my larvae have recently achieved (I’m so proud of them), I’m devoting this week’s post to describing their developmental milestones. Sea urchin larvae (Strongylocentrotus purpuratus) The pluteus (sea urchin larval form) morphology has transformed dramatically in the past few weeks. In the first picture we see the pluteus in its late prism stage, when it had recently formed its mouth and gut, and had yet to form its larval arms. In the second picture we see that the pluteus has successfully begun the development of its post-oral arms. Stay tuned--in upcoming weeks we will see posterodorsal arms developing from the anterior of its body. Sea star larvae (Patiria miniata) While at first glance the bipinnaria (sea star larval form) may not appear to have undergone significant development in the past few weeks, a closer look shows otherwise. Note how the ciliated band (the oval, outer form of the body) has lengthened and become more curvaceous. By increasing the length of this band, the bipinnarial now has more cilia (cells with whip-like tails called flagellae) to propel food particles into its mouth, aided by its two lengthening pre-oral lobes (the tip of the pointed “hat” on the anterior end of its body). Given that in both pictures the larval stomachs are filled with red algae, it is clear that my bipinnariae are successfully eating. In the first picture we also see evidence (the green substance) that my larvae can successfully defecate! Phoronid larvae (Phoronopsis harmeri) In my opinion, the antinotrochs (phonorid larvae) are the cutest of all my larvae (I know a parent shouldn’t have favorites but just look at them waving around their newly-developed tentacle crowns!). I am greatly anticipating observing morphological changes in the antinotrochs. As they continue to develop, they will eventually secrete a tube which allows them to camouflage in the mudflat sediment they reside in.
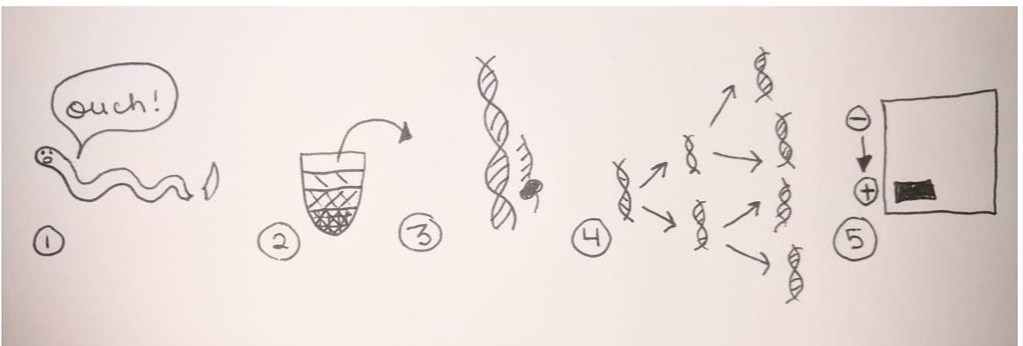
Looking at these developing larvae, I find it incredible to think that someday these larvae will metamorphically develop into their adult forms as sea urchins, sea stars, and phoronids (a type of worm). Given the dramatic morphological changes marine invertebrates make as they develop from larvae to adults, it is easy to imagine the confusion (and misidentification) past marine biologists faced in identifying organisms at different life stages as members of the same species. Next week, I will elaborate on this issue and discuss how DNA sequencing has dramatically increased scientists’ ability to identify species (and how DNA sequencing has led to the discovery of an incredible number of new species). Hello readers! Given that much of my upcoming research will involve the analysis of DNA sequences extracted from unidentified ribbon worms, I’m devoting this blog post to describing the procedures involved in procuring a specific gene of interest from a live specimen. Bear with me—this is an information-packed post—and enjoy my hand-drawn diagram of the processes involved (see below). The procedure for procuring a gene of interest begins with tissue extraction from the specimen. We obtained tissue by snipping off the tip of a live ribbon worm with a razor blade (don’t worry, the tail will grow back!). Next, we ground the tissue in a small tube with a pestle, and added detergents to break open the individual cells and liberate the carefully-packaged DNA. We then spun this mixture at a high speed until the cell components (cell membrane, organelles, DNA) had separated based on particle size and density, allowing us to extract the DNA from the uppermost layer (see steps 1-2 in the diagram below). With the DNA successfully extracted from the tissue, the next goal was to make copious copies of the gene of interest so as to create an extensive stock of the isolated gene to send for sequencing. Gene amplification is performed via PCR (polymerase chain reaction), where enzymes specific to the focal gene target and initiate the repeated copying of the gene of interest. The result of PCR is an abundance of copies of the single, isolated gene; this result is then verified via gel electrophoresis to ensure that the collection of gene copies is uncontaminated (it is easy to sneeze and accidentally amplify your own DNA!). Steps 3-4 in the diagram below illustrate the process of PCR gene amplification. Gel electrophoresis serves as a means of separating DNA fragments on the basis of size and charge. A small amount of the PCR product (copies of the isolated gene) is placed on an agarose gel which is then treated with an electrical current. The DNA migrates towards the positively-charged end of the gel; small DNA fragments migrate most quickly through the pores of the gel and create a gradient of fragment on the basis of size along the length of the gel. The resulting gradient can be visualized under UV light. For PCR product, the goal is to produce a single bright band (meaning that the DNA fragments are the same length and are therefore the single isolated product) rather than a gradient of assorted fragment lengths (see step 5 in the diagram below). Once gel electrophoresis has verified that PCR has amplified a single gene, the gene product is further purified, quantified, then shipped off to a company for sequencing. I will devote future blog entries to the exciting processes involved in determining specimen identity based on gene sequences, and will leave you this week with pictures of the sea star and sea urchin larvae we fertilized in past weeks (see below). Patiria miniata sea star larva (top) and Strongylocentrotus purpuratus sea urchin larva (bottom), imaged under a compound microscope.
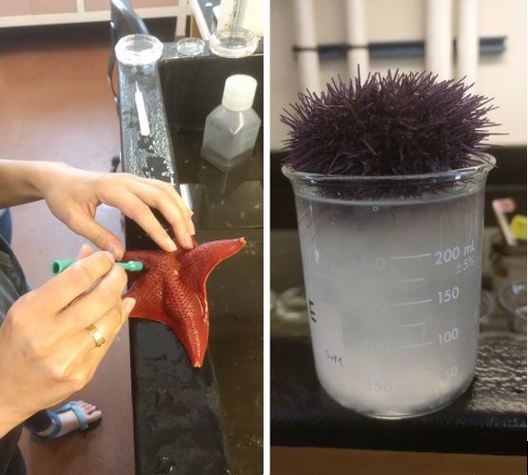
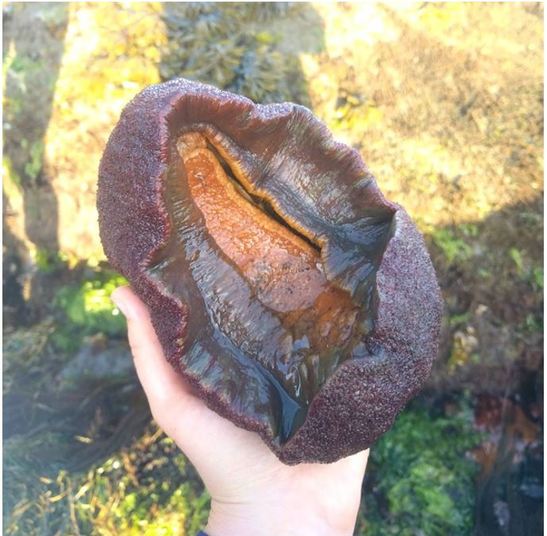 The OIMB community is filled with an enthusiasm for animal biology unlike I’ve ever experienced before. Here, conversations from lab spill over into lunch and I am endlessly amazed by the depth of knowledge I encounter in my peers. During our conversations, I often wish I could live a thousand lives each devoted to the intense study of a different organism; I want to study deep sea larval activity and barnacle parasitism and chiton larval morphology, and I feel incredibly fortunate to be surrounded by amazing researchers whom I can live through vicariously. This is the contagious, effervescent energy of OIMB, where professors and graduate students alike welcome you into their labs to observe larval cultures, crab tanks, and collections of comb jellies dancing in their jars. While there is much to learn at the lunch table, there is even more to learn by surveying the local ecosystems. My first morning of tide pooling, I felt as if I were exploring an alien planet. We arrived at the beach in the early morning, when the tide was very low, to a vast landscape of algae and kelp-covered rocks which jutted at all angles to form an array of tide pools. As I clambered about, I stepped on something squishy that I assumed to be a rock—and met my first gumboot chiton (see picture top left). Nicknamed the “wandering meatloaf” for its reddish-brown hue and fleshy body, Cryptochiton stelleri are common to local tidepools (and easy to mistake for rocks!).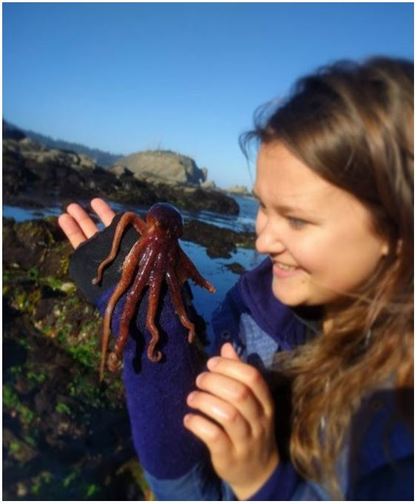  The contents of the tidepools themselves were equally bizarre and enthralling. While anemones, sea urchins, and sea stars resided in many tidepools, no two tidepools were exactly alike. Fish darted about in some, while colorfully-spotted nudibranchs undulated across the bottom of others. I think that the most exciting find of the day, however, was this small octopus! Although I encounter fewer octopuses in the lab, I find my work inside just as enthralling as tide pooling. My training for Svetlana’s lab has focused primarily on practicing lab techniques for extracting DNA, amplifying specific genes, utilizing gel electrophoresis to verify the successful amplification of the specified genes, then purifying and quantifying the isolated genes before sending them off for gene sequencing. These lab techniques will enable us to utilize DNA barcoding to determine new or cryptic (morphologically indistinguishable but distinctly different) ribbon worm species. Much of Svetlana’s work centers around the nuances of determining what a biological species is, and I am excited to delve more into this complex subject. My training for George’s lab has focused primarily on sea star (Patiria miniata) and sea urchin (Strongylocentrotus purpuratus) fertilization, oocyte injection, and familiarizing myself with compound and dissecting microscopes. This week my fellow lab interns and I learned how to extract sperm and eggs from both sea stars and sea urchins, and had the opportunity to watch the fertilizations occur live under a compound microscope. To extract sea star gonads, we utilized a biopsy punch to make a circular incision on the side of a sea star arm and then removed the gonads from the body cavity using a pair of tweezers. To extract the sea urchin gonads, we vigorously shook the urchin for several minutes until the distressed specimen expelled a cloud of eggs/sperm from its gonopores into a beaker of seawater (see pictures below). We’ve been watching our fertilized oocytes develop into larvae, and have even observed the blastula formation of developing sea stars using a time lapse video (check out the link below). I promise to post more pictures of the larvae as they develop.
https://youtu.be/tLQd79pH3VA My brain is overflowing with new information, and I hope yours is too. Next week I’ll delve more into describing my research projects—until then, take care, and comment below if you have questions! |
Author“Instructions for living a life./ Pay attention./ Be astonished./ Tell about it.” (Mary Oliver) Archives |




 RSS Feed
RSS Feed