|
This has been our last week here for the summer and we spent it finishing the guide, printing the guide, and presenting our posters. We have been extremely busy but it has been so exciting to see our work from the last nine weeks come together into our final products. We processed 120 individual worms and obtained DNA sequences for 92 individuals. We found 36 species in total, 22 of which were hoplonemerteans, and each received a card in the identification guide. We found four nemertean species that are new to science, two species that have not been found in Oregon before, and an adult individual of a species that has not been found in the adult form before. It has taken an immense amount of work to get this far, but it has been entirely worth it and I have learned so much in the process of making our identification guide.
I want to use the rest of my final blog post to thank everyone that helped make this one of the most amazing summers of my life. I have been genuinely happy here and am utterly grateful for this experience. Svetlana has been a truly amazing mentor and has taught me so much about nemerteans, marine biology, and science in general. I am so incredibly thankful to have spent my summer working in her lab and hope to follow in her footsteps to become a researcher one day. It has also been a privilege and a pleasure to work with Christina, the graduate student in our lab, who helped me work through the little kinks of research I did not see coming and readily welcomed me into the lab. Likewise, I have loved working with Frances Conable, an undergraduate in the lab, in a variety of situations throughout the summer. Additionally, Rebecca, my fellow lab mate, helped the many hours we spent in the lab fly by. Even the tedious and stressful aspects of worm dissection and photography were fun thanks to her. Although science may be perceived as dry and sterile, the people I worked with this summer have been integral in shaping my experience in research and contributed to it for the best. I want to thank Richard Emlet and Maya Wolf-Watts for running this wonderful REU program and helping us edit and revise our posters, as well as Nicole Nakata for acting as an impeccable REU den mother throughout the program. I also want to give a shout out to my fellow interns, who have been some of the nerdiest, funniest, and most enjoyable people I have ever met. My summer would not have been the same without them. Finally, I want to acknowledge my family and friends for their endless support, as well as my blog readers for learning about my work. After I leave OIMB, I will return to my home institution and continue pursuing biology to eventually attend graduate school and earn a PhD. This program has solidified my resolve to pursue a career in scientific research and has exposed me to marine biology, which has become a particularly interesting aspect of biology to me. I would not be surprised if I end up studying marine biology for the rest of my life; but whatever I do with my career, I know for sure that I will look back on my time here at OIMB fondly and as a fundamental step in my path to becoming a professional scientist.
0 Comments
We finished all of the molecular work this week, it will be all computer work from here on out. We ran 109 PCRs (polymerase chain reactions that amplify DNA) and ended up sending 44 of them for sequencing. Now we can use these last sequences in our guide.
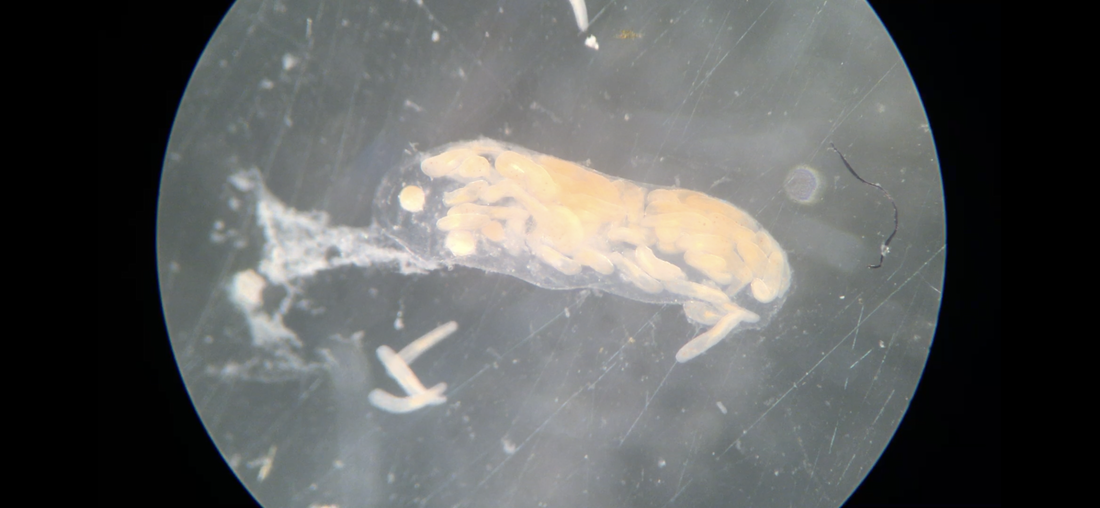
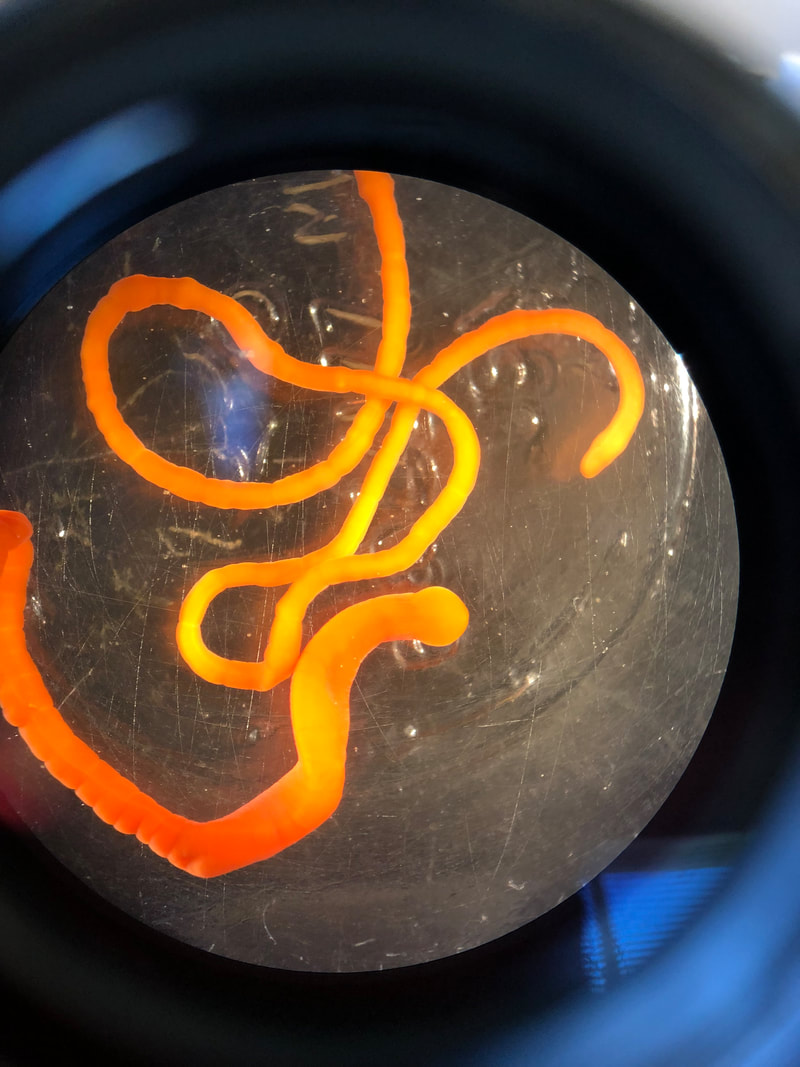
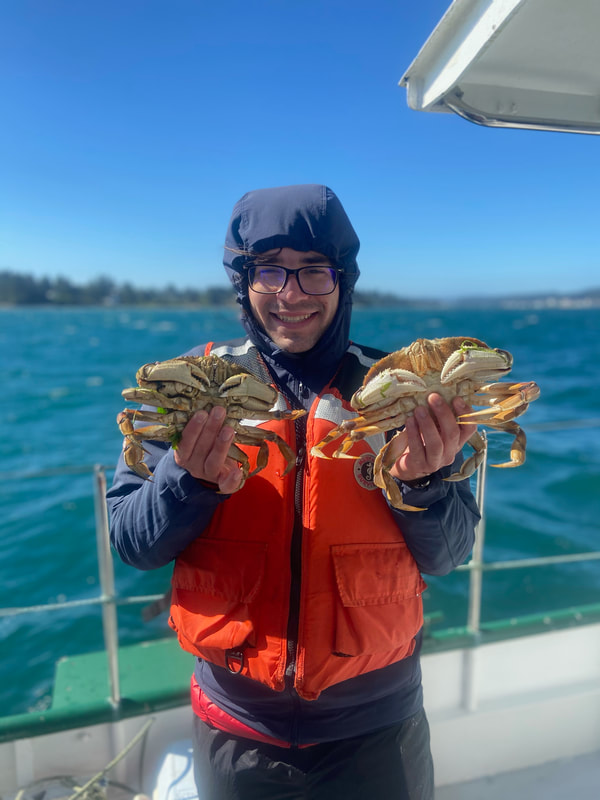
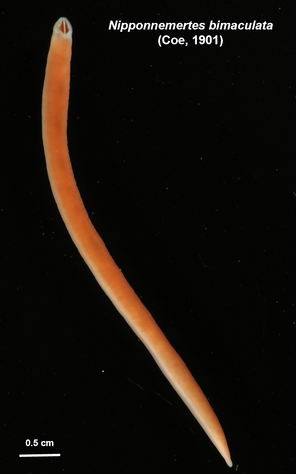
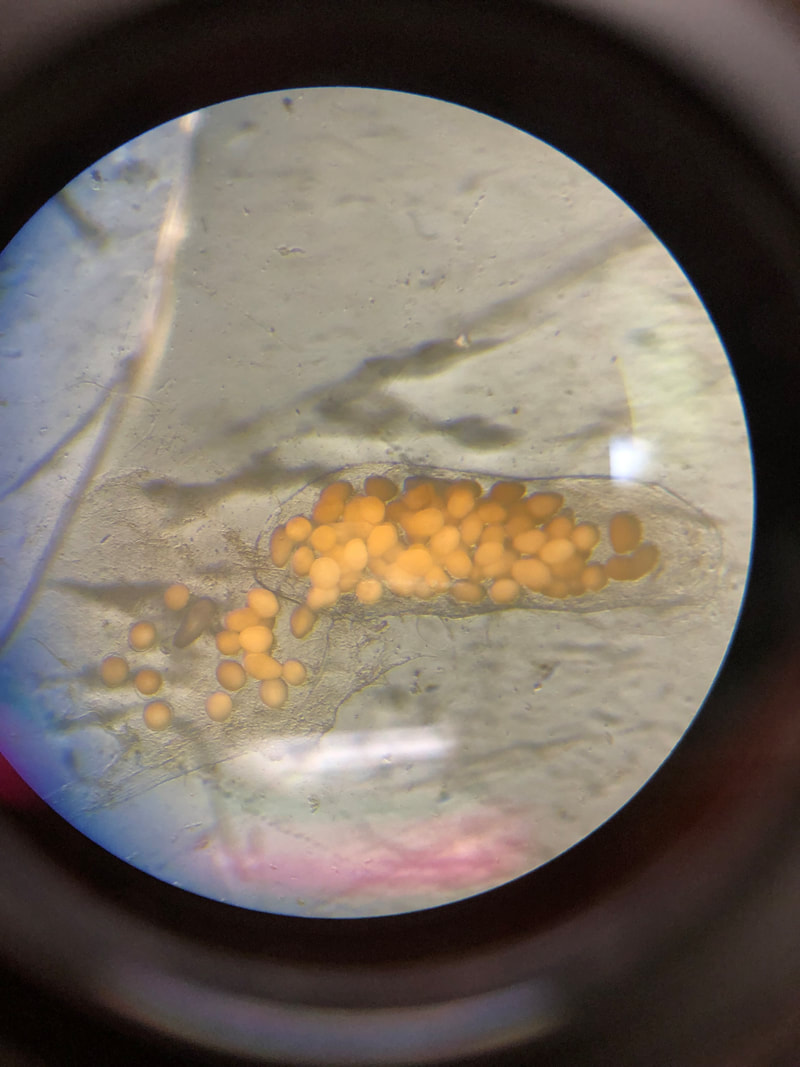
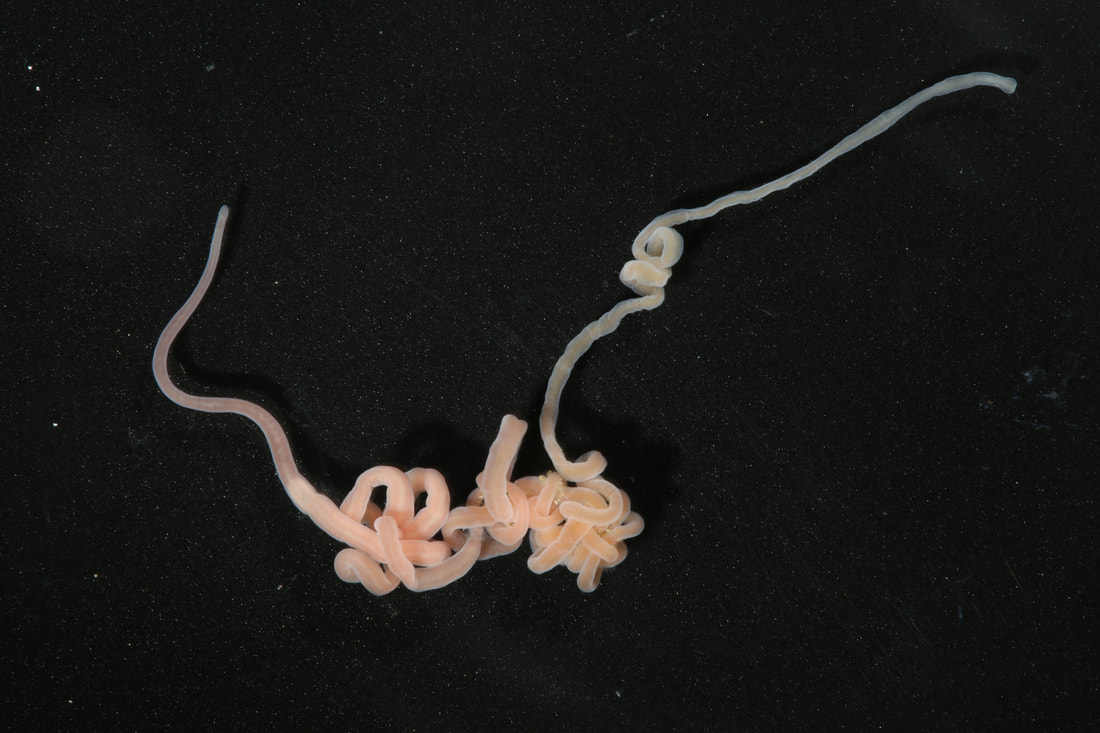
We have started to put the guide together and sneaking a peek of our final product has been really exciting for all of us. The guide will act like a flip book that a worm enthusiast could carry in their pocket when they are out in the field, with one double-sided page for each nemertean species. There are two aspects of the guide that set it apart from existing guides. The first is that each card has a large color photo of a specimen of that species, as existing guides contain only black and white photos or line drawings of individuals. The picture entirely takes up the front of each card. There will be information about the species range and habitat on the back, in addition to a picture of the central stylet for hoplonemerteans. The second crucial aspect of our guide is that species identifications are verified with DNA barcoding so that we know each photo in the guide actually belongs to the species it represents. This is why we have spent so much time and energy on the molecular side of the project and working with sequences on the computer. All of the interns will be presenting our research projects at a poster session held on the last day of the program, which is next week. As the more labor intensive parts of my project have quieted down, I have been spending a lot of time designing and making my poster so that it can easily be understood by someone who is potentially unfamiliar with nemerteans and biodiversity surveys. It is definitely a challenge to balance brevity and comprehensiveness when talking about my project. I also did not realize ahead of time how logistically difficult it is to fit everything I want to talk about onto a 1,400 square inch (which is a lot less than it sounds) poster. Waiting on the last of our results to complete the poster reminds me how close we are to the end of the program and how soon we will be leaving the field station, which is a little sad. However, I am excited to present the poster to people who are (hopefully) genuinely interested in what I worked on this summer at the poster session. Below is the species card (front and back) for Nipponnemertes bimaculata, one of my favorite ribbon worms. There is a genus of ribbon worms, called Carcinonemertes, that lives on crabs and eats their eggs. This type of worm is known to occur on species such as Dungeness crabs, an economically important species since they are fished for human consumption. This week, some female striped shore crabs came into our possession and we saw that they were brooding (carrying eggs on their bodies) so we decided to check the eggs for Carcinonemertes. Previously, Carcinonemertes has not been shown to occur on striped shore crabs, so we weren’t sure if we would find any. As I was pulling the eggs off the crab, I found a Carcinonemertes nestled among them. What is even more exciting is that the worm had eggs in it that she eventually released into the bowl in an egg mass (however, we probably won’t get baby worms from them since they weren’t fertilized). If our preliminary identification of the worm is confirmed by DNA barcoding (which we are currently working on), we will be able to say that Carcinonemertes occurs on striped shore crabs and improve our understanding of the role Carcinonemertes plays in the ecosystem and what crab species the genus interacts with. We never planned to sample crabs for worms while making our identification guide, but this was an opportunistic find that ended up being very exciting for us. We finished photographing and extracting DNA from the last of our worms this week. I feel immensely more comfortable with the entire process of documenting worms and using a variety of chemicals to extract DNA from them, when compared to my confidence levels during my first week here. It was very bittersweet to realize we will not be doing this again before the REU program ends. However, I also noticed how far I have come in the relatively short time I have been here, which will give me momentum as I continue pursuing a career in science. On a more random note, I fertilized some sea star eggs my first week here at the Oregon Institute of Marine Biology. I decided to check on them this week and saw that one of the larvae has finally settled (moved out of the planktonic stage that swims around in the water and attached itself to the bowl it is in while looking like a sea star)! This week we finally began working on the DNA sequences we got back. When we visualize the sequences, we can basically read the genetic code for that worm (a small portion of the code might read something like: C A G A T T A). However, sometimes the code gets cluttered and there are two letters in one place (so the same small portion of the code might read something like C A G/C A T T A). We have to go into the code and take out the letter that should not be there (which is the extra C in this case). We have two copies of each sequence so we can use them to proofread each other; that is how we know which letters should and should not be there. We also have to trim off the ends of the sequences since they are often really messy or do not tell us anything. Luckily, most of our sequences were not very cluttered and we cleaned them very quickly. We have run all of our sequences through a database but have only had time to look at a few of the matches we got. However, we have already confirmed one species ribbon worm not previously shown to live in Oregon does, in fact, occur here. This is exciting because the same species has been found in California and Washington, meaning that we can show that their range is at least somewhat continuous along the coastline. We spent one morning this week out collecting more worms at the boat docks near the lab. We are hoping to get worm species we haven’t found in the rocky intertidal zone or the mudflats yet. We are also planning to obtain some worms in kelp holdfasts (the root-like structure that kelp uses to anchor itself to the sea floor) through SCUBA diving later this week, weather permitting. This will give us even more species of worms we haven’t seen in the places we have already sampled. These two collections will be the last ones we do before the program ends but will give us plenty of material to keep us busy for the next few weeks. Although I am studying marine biodiversity for the REU program, I had an amazing opportunity to help put in a new forest research plot in the South Slough National Estuarine Research Reserve with Dr. Alice Yeates and Anna Liang over the weekend. They are studying different areas of the forest to help inform management decisions in a local estuary (the area around where fresh water from rivers meets salt water from an ocean). I have never done any sort of forest research before so I learned a ton of new skills, such as: assigning trees ID numbers, measuring tree height, and finding diameter. It was fantastic to get out and explore beyond the coast; hopefully, I will be able to go out again with them soon. Although we have gotten our first sequences back, we have not started to clean them up and run them through a database full of sequences from across the animal kingdom (and beyond). This is because we are so busy processing new worms! We have spent three mornings collecting nemerteans from mudflats (a habitat I have not been in before) this week and have loads of worms to document and preserve. This time around, we found different worms from the ones we found in the rocky intertidal when we spent some time collecting in that region. Many of the worms from the mudflats are in the classes Pilidiophora and Paleonemertea, so they will not be going into my identification guide (mine will cover the remaining class, Hoplonemertea), but will be in Rebecca’s guide. However, we found some Hoplonemerteans so I processed stylets for those species and found that some individuals of the same “species” (really a cryptic species complex) have a different number of accessory stylet pouches (where backup stylets are made and stored), possibly providing a way to differentiate between species in the complex. Collecting in the mudflats is very different from doing so in the rocky intertidal. Rather than scouring rocks and algae for worms, we dig in the sand using shovels. It is a lot more labor-intensive and involves a lot more luck. We basically scoop out mud and look for worms as it falls apart in the shovel. The issue is that the worms are often destroyed by the shovel blade so we have to be lucky that the blade does not come into contact with them under the sand (since we can’t see where they are before we dig). However, we did obtain some worms entirely intact and were very excited to obtain such valuable and informative specimens (having the complete worm gives us a lot more identifying features to work with than only a body fragment does). While digging for worms, many locals came up to us to ask what we were doing. Most of them thought we were digging for clams and were surprised that we were looking for worms. It was very rewarding to interact with the public and tell them about our work. Many of them were genuinely interested in our research, and it feels great to be working on something that others care about (even if only while they are talking to us). I remember talking to people that studied biology and being amazed by them so it was extremely fulfilling to be on the other side of that interaction by helping others become interested in marine animals. We have been so busy processing worms this week that we kind of forgot about the eggs I mentioned in my last post. However, we were lucky enough to check on the egg masses right as the baby worms were exiting them! I put a picture down below. We think these are a species in the genus Tetrastemma (a group of Hoplonemerteans). Some tetrastemma are known to lay their egg masses on surf grass, although this is not very common in ribbon worms. We don’t know exactly how long it took the tiny worms to emerge since we don’t know when the masses were fertilized, but it took no longer than a month for them to develop from eggs into mobile worms. 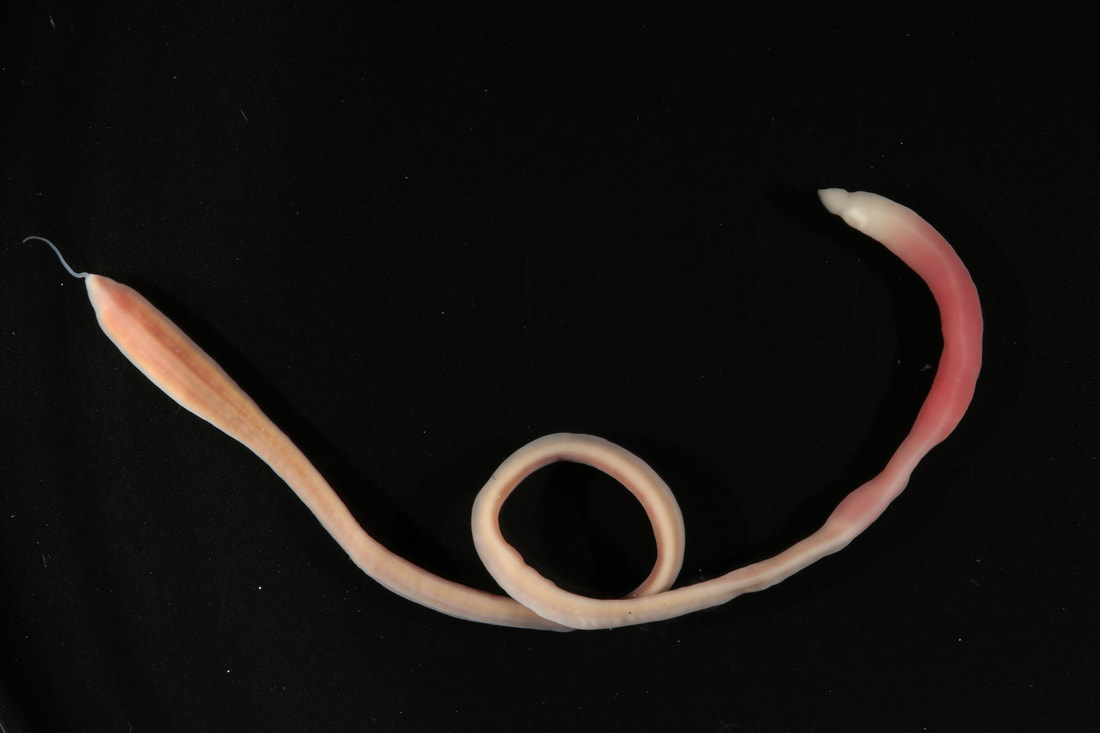 This is a Maculaura magna, a worm we found in the mudflats. Despite spending most of its time buried in mud, the front part of its body is a vibrant pink. These worms have no eyes and use something more similar to our sense of smell to get around the mudflats. The little thread on the end of the worm is not technically considered a tail but is called a “caudal cirrus”. Like many other aspects of nemertean biology, no one knows what it is used for. I have spent a lot of time in the lab this week working on the molecular side of my project. Even though I am in the lab from morning until dinner extracting DNA, amplifying it, and purifying it, the time goes by extremely quickly. I suppose this is what happens when you enjoy what you are doing, and I am extremely happy doing this work. And although I am done with working with the DNA directly for a while, I will come back to it once we collect more samples from the field.
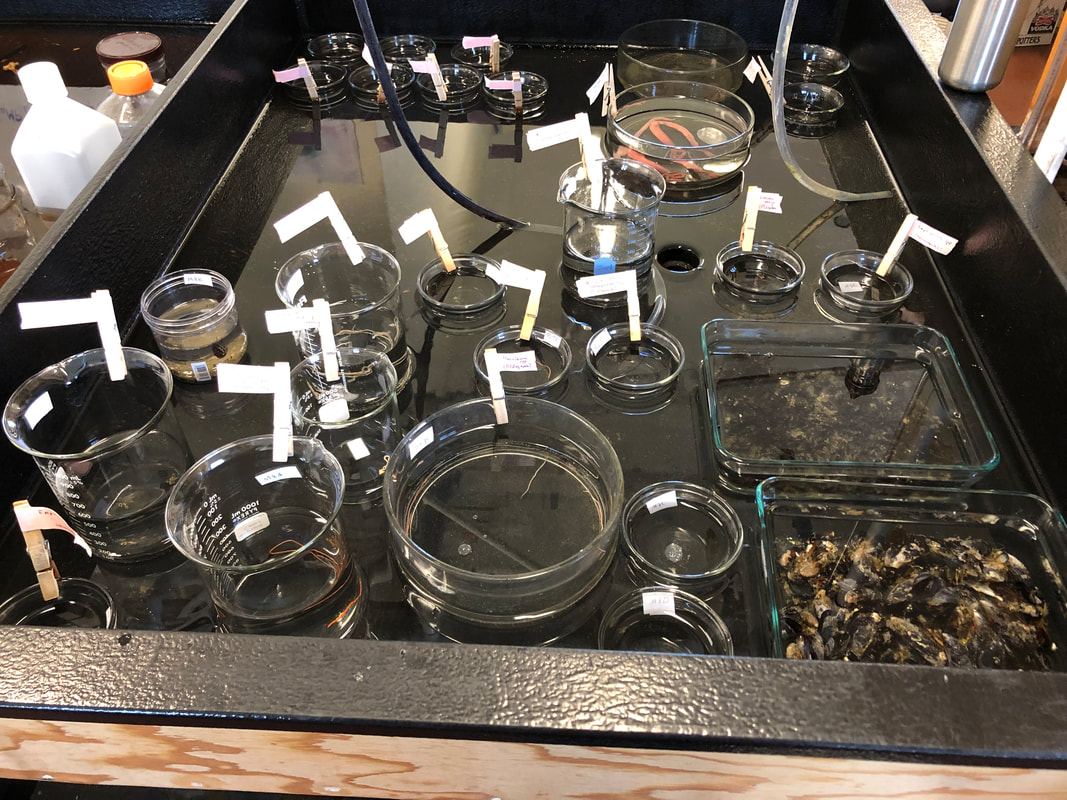
While changing the water in the bowls we keep the nemerteans in, one of my lab mates found some egg masses laid on the glass. This was quite the surprise, since most nemertean species release their gametes (eggs and sperm) into the water and fertilization occurs rather randomly (not all in one place like we saw in the egg masses). There are about four clumps of eggs in the bowl, which means we will hopefully get to watch worms hatch out of them! We may also gain insight into what the eggs look like as they mature, so this will be both fun and informative for us. Working with live animals can sometimes be frustrating (such as them escaping) but there are also moments like this when something completely unexpected happens and that is something I am starting to appreciate about biology. I am looking forward to learning more skills next week, as we just sent our first round of DNA samples for sequencing. Once we get the sequences back, I will practice cleaning them up (DNA from each specimen is sequenced twice so we have to fix any discrepancies between the two sequences we get) and run them through a database to compare with previously published sequences. Getting the sequences back and analyzing them is super exciting for me because I will finally get to see the fruits of my labor working with the DNA. Until now, I have only seen liquids of various colors containing DNA (the DNA itself is too small to see with the naked eye) and indicators that DNA is present in the solution. Soon, I will have a more tangible product that I can read and use to make meaningful comparisons between individual worms. I will also have trustworthy identifications of the ribbon worms we sequenced, ensuring that the identification guide I make will be accurate. We plan to collect nemerteans in a new type of habitat, mudflats, in the coming days. We hope to find species different from the ones we found in the tide pools of the rocky intertidal zone in past weeks. Once we have collected our new worms, we will start the entire process over again: photographing, making vouchers, and DNA analysis. Updates are to follow. After collecting more worms during extremely low tides, we have spent a lot of time in the lab processing them. We took an assembly-line approach to photographing, taking sections of worms for DNA analysis, and preserving the head/front half of worms. Photographing the worms takes both skill and luck on our part and cooperation on the worm’s part. We have to be sure that the lighting is just right and that the camera is focused so that the worm’s few identifying features are sharp. No matter how great the preparation is, we still need the worm to hold still, which is not something we can control. However frustrating this can be, seeing the images on the computer makes it worth the effort. Without these images, our identification guide would be vastly inadequate by failing to show the visual stunningness and very minute details of worms that can be used to tell species/complexes apart. I have spent most of my time this week cutting bits of the posterior (back) end of worms off for DNA analysis. After cutting each section off, I put it into alcohol and then into a freezer so that we can extract DNA from the preserved tissue in the coming weeks. I also spent some time preserving the other (anterior, or front) part of each worm into formalin, a fixative we use to keep the tissue from degrading. We then store this portion of the worm in a museum in case anyone ever wants to look at them and their insides. Many of these morphological vouchers (the preserved front part of a worm) came out very well, which was very pleasing for me. Since I will be working with the class Hoplonemertea, I will examine the stylets (a dagger Nemerteans use to inject venom into prey) of various species and cryptic species complexes. Hence, I have been extracting the stylet-region of the proboscis (what they use to launch the stylet into their prey) from worms this week. Like many other aspects of working with Nemerteans, this part is conducted under a microscope. I have spent more time using microscopes this week than I ever have before and am becoming very comfortable with them. I thought that extracting the proboscis would be easy after watching Svetlana (my mentor) do it effortlessly, but there is definitely a learning curve to it. But patience and perseverance have paid off, as I am now able to extract the stylet-region of the proboscis intact. When I placed my first stylet under the microscope and saw that it was photograph-ready, I found Rebecca (my lab partner) and told her because I was so excited. I have begun to realize that no matter how straightforward I think something will be, it can easily become more complex. Before preserving one of the exceptionally long worms, I had to untie the knots in it under a microscope for about 20 minutes. Typically we can just put the worm in a chemical that preserves it, so this extra step was definitely unexpected. I did not plan to untie a worm with forceps when I woke up that day but it was very satisfying to successfully untie the worm without breaking it. Such is the life of a Nemertean scientist. We will continue processing worms in the next week and begin circling back to the molecular side of the project soon. I will get a lot more practice preserving worms and looking at their stylets next week, so I will be a pro by the end of the program. Updates will be forthcoming. I have spent my first two weeks at OIMB planning and preparing for my project. For which, I will be creating an updated identification guide to Nemerteans (also known as ribbon worms) found locally around Coos Bay. As of now, existing identification guides for ribbon worms are vastly inadequate for a multitude of reasons. Firstly, many Nemertean species are undescribed, some of which are cryptic and look like other species. Secondly, many species are called by the wrong name, being mislabeled as another species that inhabits a completely different geographical area. For example, a local worm was inappropriately given the same name as a European Nemertean that it looked similar to, despite being a completely distinct species. Additionally, one species may be listed under multiple names due to intraspecific variation, in which individuals of the same species may look vastly different from each other. Thirdly and finally, while existing guides provide photographs of worms, they are not in color. As Nemerteans often have striking colors or very few features, it is important to show the color and shape of them to make identification easier and more accurate. Therefore, my identification guide will be a picture guide to the most common and most conspicuous species of intertidal ribbon worms in Southern Oregon that corrects the names of inappropriately named species and alerts users to the presence of multiple cryptic species (ones that look very similar to each other) when they occur.
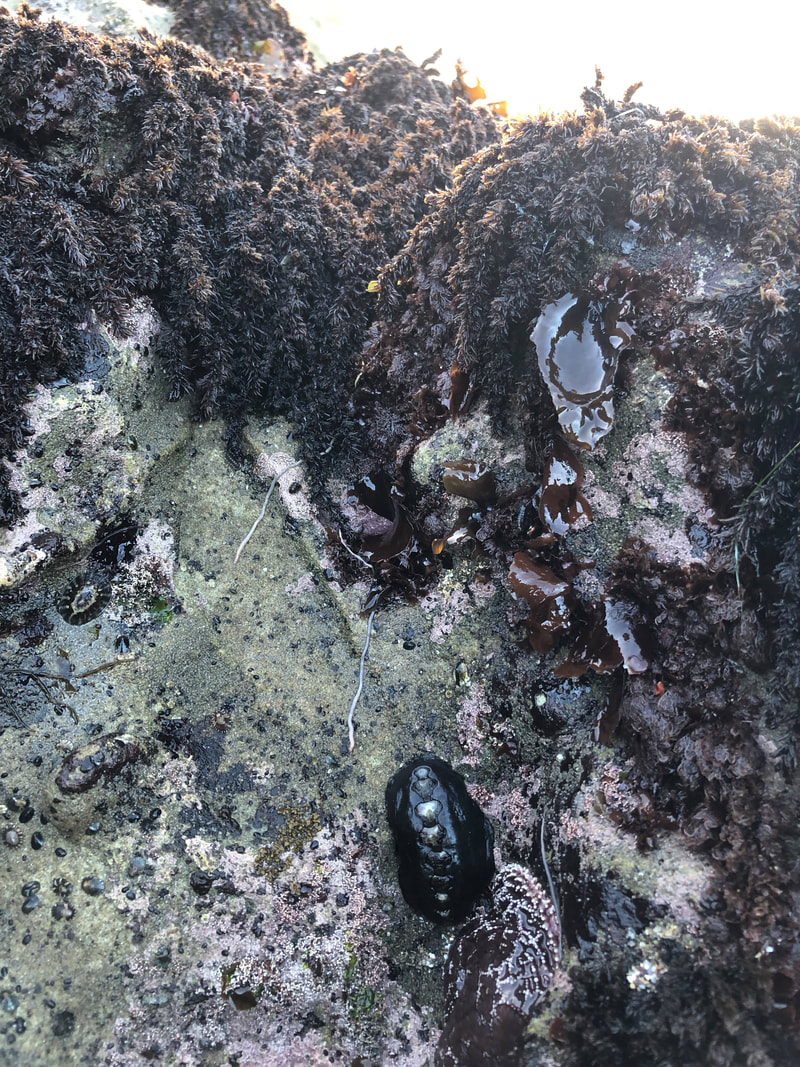
Specifically, I will focus on the class Hoplonemertea, or ribbon worms that have a stylet, a dagger-like structure that Nemerteans use to stab prey with while hunting. I chose to work on this group since they have such a unique mode of predation and it will be a challenge to examine these stylets under a microscope. To begin the process of creating the identification guide, we capitalized on the exceptionally low tides this week to collect worms in the intertidal zone. The worms were exceptionally difficult to spot at first (being only a few millimeters or centimeters wide), but I found a few by the end of the field trip. This was my first “professional” tide pooling expedition and was very informative for me in regards to both Nemerteans and other marine invertebrates. On the molecular side of the project, I learned how to extract DNA from worms, amplify it, and purify it. For this project we will be looking at the cytochrome c oxidase subunit one gene, or the COI gene. We use this gene because it mutates quickly, so that each species has a unique code in this section of their DNA. Once we obtain the isolated genetic sequence, we will send it to a lab for sequencing. Upon receiving the sequence, we will clean it up and run it through a database. When comparing our gene to other COI genes in the database, we want to determine whether the species has already been described or not, if the species name is not meaningful (the worm has been grouped with species not related to it), and if there is a cryptic species complex (a group of worms that look alike but represent distinct, often closely related species). Once we have this updated information about locally found ribbon worms, we will begin writing the identification guide. It took me an entire week to learn the molecular techniques that I will be using this summer, as I have never worked with DNA in this capacity before, but I am happy to be learning such a scientifically universal skill and I already feel way more comfortable in the lab than when I started. I am still working on identifying the worms using morphological features (ones we can see with a microscope) prior to our genetic analysis of them and there is a recurring theme of Nemerteans looking extremely, almost ridiculously, similar to each other. It will be a challenge to differentiate between “cream-colored featureless worms,” but I am excited to continue learning about such an understudied group of marine invertebrates. In the coming days, we will begin processing our worms by prepping them for DNA analysis and photographing them for the identification guide. Updates on that will be coming soon. |
AuthorI am Jacob and am from Rancho Cucamonga, CA. I am a rising Junior at Pomona College in Claremont, CA who has never studied marine biology in school before but has always been interested in it. I am incredibly excited to spend the next two months working in Svetlana Maslakova’s lab studying Nemertean (ribbon worm) biodiversity through genetic analysis. Archives
August 2021
Categories |
Proudly powered by Weebly












 RSS Feed
RSS Feed