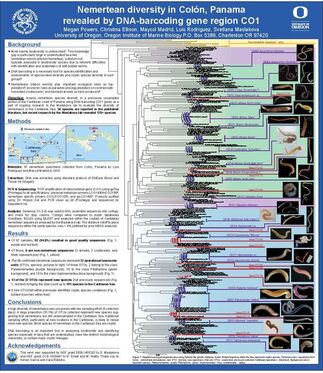
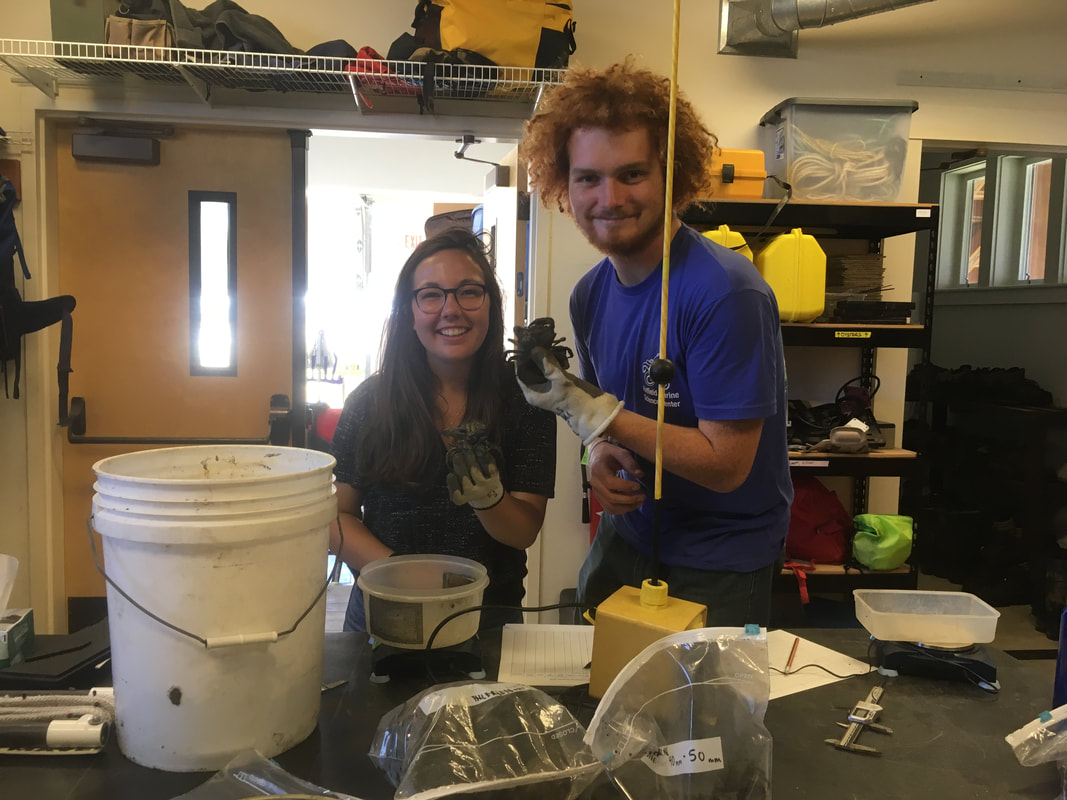
 Nine weeks, 97 worms, and 13 new species later, my time here at OIMB has come to a close. This week we finished our posters, and presented them to the OIMB community. We had a great turnout so we were able to present our research many times throughout the poster session. I am going to miss a lot of different aspects of OIMB. First of all, I will miss the research. This summer I realized how much enjoy taxonomy and systematics. This realization will help me to better choose a graduate school as I move forward in my research career. I am also going to miss Oregon (the ocean in particular). This was my first summer of tidepooling and hiking through the woods to hidden beaches, and I never want to go back to good old landlocked Iowa. I have no doubt that I will be back to the Pacific Northwest in the near future! Most importantly though, I will miss the people. Everyone from my mentor and the staff at OIMB to the University of Oregon students and my fellow REU interns has made an important impact on my life. During the final week we spent as much time as we could together hiking, measuring crabs, and hanging out on the beach. I know that through this experience I have done some amazing research and made lifelong friends. I am so grateful to have had this opportunity.
0 Comments
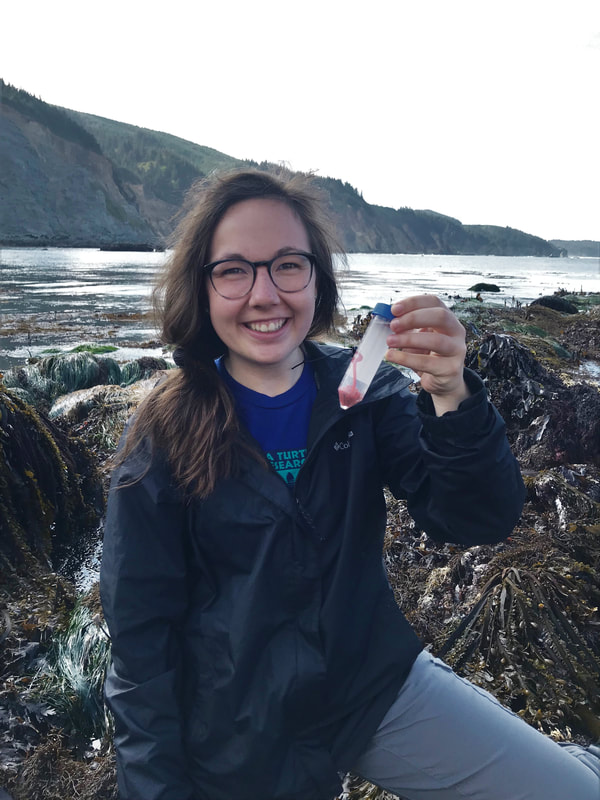
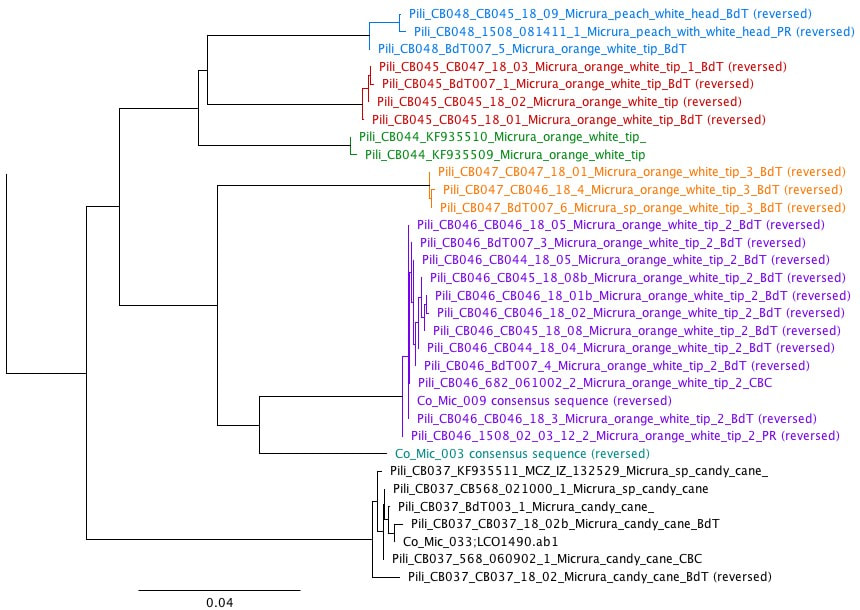
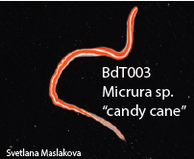
 Eight weeks of work have come to an end, and we are working summarize a summer of work in a poster. That is a lot to fit in a 35x40 inch space, especially when you have to fit a huge phylogenetic tree and pictures of all of the worms that we worked on. I am currently taking a break from inching photos across a computer screen one by one to write this blog post. While the process of making a scientific poster is a bit long and frustrating, we will be able to take our posters home to keep and (hopefully) present at conferences or undergraduate poster sessions. It is also rewarding to see all of your hard work in put together at the end of a project. Last Saturday, we had the chance to practice communicating our research to the general public. Each lab set up a table at the Charleston Marine Life Center, a museum/aquarium across the street from campus. Adrian and I brought a few different local species of nemerteans to show visitors. We talked about the what ribbon worms are and their importance, but we also discussed marine biodiversity in general. The majority of marine species have not yet been described, and that is part of what makes our work to discover new worm species so important. We need to know what is out there if we want to understand how each animal contributes to an ecosystem, and how we can protect it. Another benefit of learning what is in our oceans is that we may find something useful. Nemerteans, for example, produce mucus that is toxic to their prey, but those toxins have been studied for potential medical uses. It was fun to see that people are interested in the research we are doing here. At the end of last week, we visited a new tidepooling location called Qochyax Island. The tide was very low and we found all sorts of worms, algae, isopods, and other organisms. The tidepools were teeming with life. This week I sent my last batch of worm DNA samples away for sequencing. Typically, we get our sequences back just two days later which is becoming more and more helpful as our time here draws to a close. We are now starting to compile our data into what is known as a phylogenetic tree. A phylogenetic tree is a way to visualize how different groups of organisms are related. In the case of my research project, I am making a phylogenetic tree with all the CO1 DNA barcoding sequences for specimens that I have been working with from Colon along with all the nemertean CO1 sequences that Dr. Maslakova has previously collected from the Caribbean. Including all the available nemertean DNA sequences from the Caribbean allows us to compare our samples to those that have already been found. The lines leading up to each specimen name represent the genetic difference between the CO1 genes of any two specimens. The longer the lines, the longer it has been since those two specimens or groups diverged from a common ancestor. The shorter the lines, the more closely related two samples are. In the tree above, there are 6 species. I knew that each of these groups were a distinct species because the lines that connect them are very short, but the lines between species (different color groups) are much longer. Specimens are considered separate species if there is more than a 4% (0.04) difference between their CO1 genes. This distance is represented by the scale bar at the bottom of the tree. The interesting part about this set of 6 species is that the 5 of the species that I have indicated with colored text all look almost exactly the same! They represent a “cryptic species complex” because the worms look the same, but are genetically 5 distinct species. You can see in their specimen names that they are all orange with a white tip. The 6th species at the bottom known as “candy cane” because it has not yet been officially described as a species has bright red and white stripes. Last weekend, about half of the REU interns had parents in town. We gave our parents a tour of the labs, campus, and the Charleston Marine Life Center. My parents and I also checked out a couple of the nearby state parks and went tide pooling!
|
AuthorHi! I’m Megan Powers, a fourth year Biology and Environmental Sciences student at the University of Iowa. Throughout my summer at OIMB, I will be working with the Maslakova lab to assess Nemertean diversity in the Caribbean. Archives
August 2019
Categories |
Proudly powered by Weebly







 RSS Feed
RSS Feed