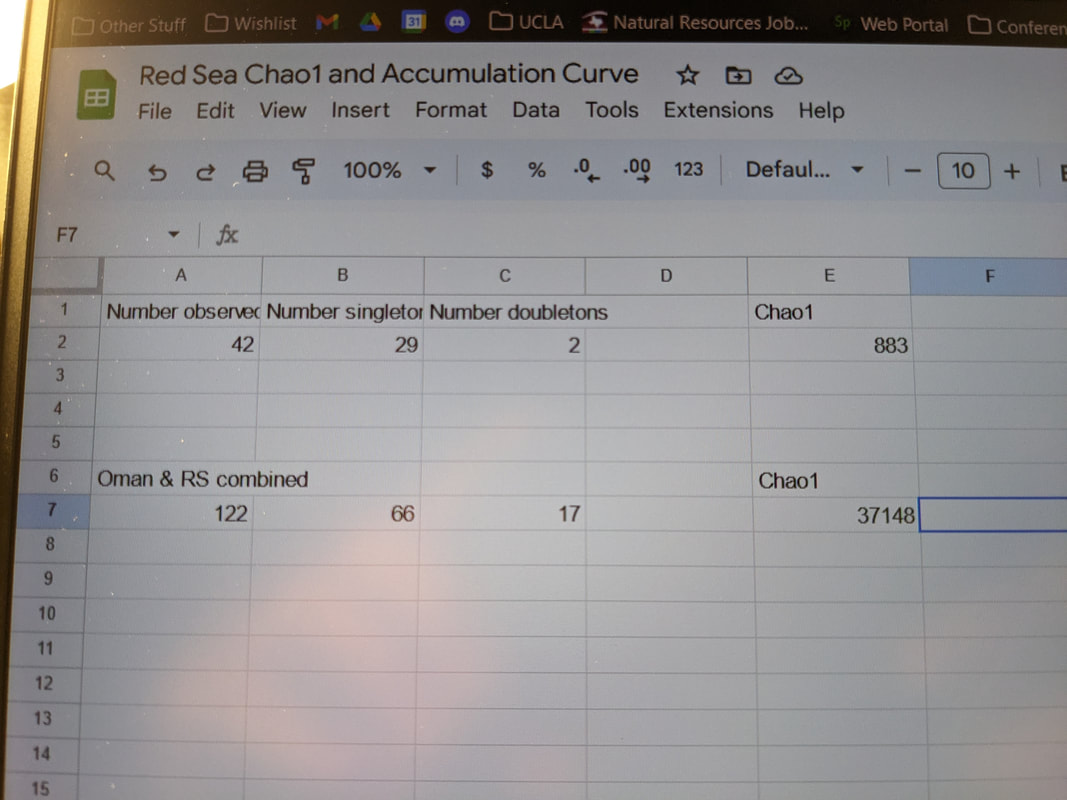
"What would the world be, once bereft Inconceivably, unerringly, indisputably, I have arrived at the end of this REU program. Our submissions of specimen records, DNA sequences, chromatogram trace files from the raw output of the sequencing process, and identifications have been uploaded to the Barcode of Life Database. My poster is printed, and as you read this, I have presented at the OIMB REU symposium and already flown home to North Carolina. This has been a deliriously wonderful summer, replete with late nights pipetting precise, minute aliquots of strange, clear liquids into other, precisely measured clear liquids. I have learned every step of the genetic sequencing process, from extraction, to PCR, to PCR, to re-PCR, to troubleshooting PCR, to one final PCR with fresh primers that works like a charm, to agarose gels, to loading plates, to cleaning and processing sequences, to making the data accessible on a public database. I found out how easily mistakes creep in, and how to root them out with meticulous records and by double-checking every step of data management and analysis. I have discovered an entire phylum which I had barely heard of before this internship, and though hardly an expert, I can catch and identify a couple of genera, and I have learned the basic taxonomic divisions of Nemertea. I have also submitted an abstract to a conference for the first time, and I am planning to present my research at the Society of Integrative and Comparative Biology's 2024 meeting in Seattle! More to the point, by my nine weeks of effort, I have documented 44 species from the Red Sea, and demonstrated that there are over a hundred more that are waiting to be discovered. Never before have I been so close to the cusp of new discoveries, nor more acutely aware that there are uncharted universes of biodiversity, in all its wriggling, crawling, squirming, sessile, chemosynthetic, carnivorous, parasitic vivacity, propelled by the simple fact of natural selection to unimaginable variety. There is so much that is known, and yet so much more lies just over the horizon, or perhaps just under the water's surface. Of course, this summer would not have been one fifth the joy that it was without the support and camaraderie of my fellow REUs. Their positive qualities are too numerous to recount, but rest assured they are each unique and wonderful, and have excellent blogs that you can peruse! I am beyond grateful for the opportunity to meet all of them and explore this pocket of Oregon together. Be happy and healthy. "Something told him that something was coming to an end. Not the world, exactly. Just the summer. There would be other summers, but there would never be one like this. Ever again.
0 Comments
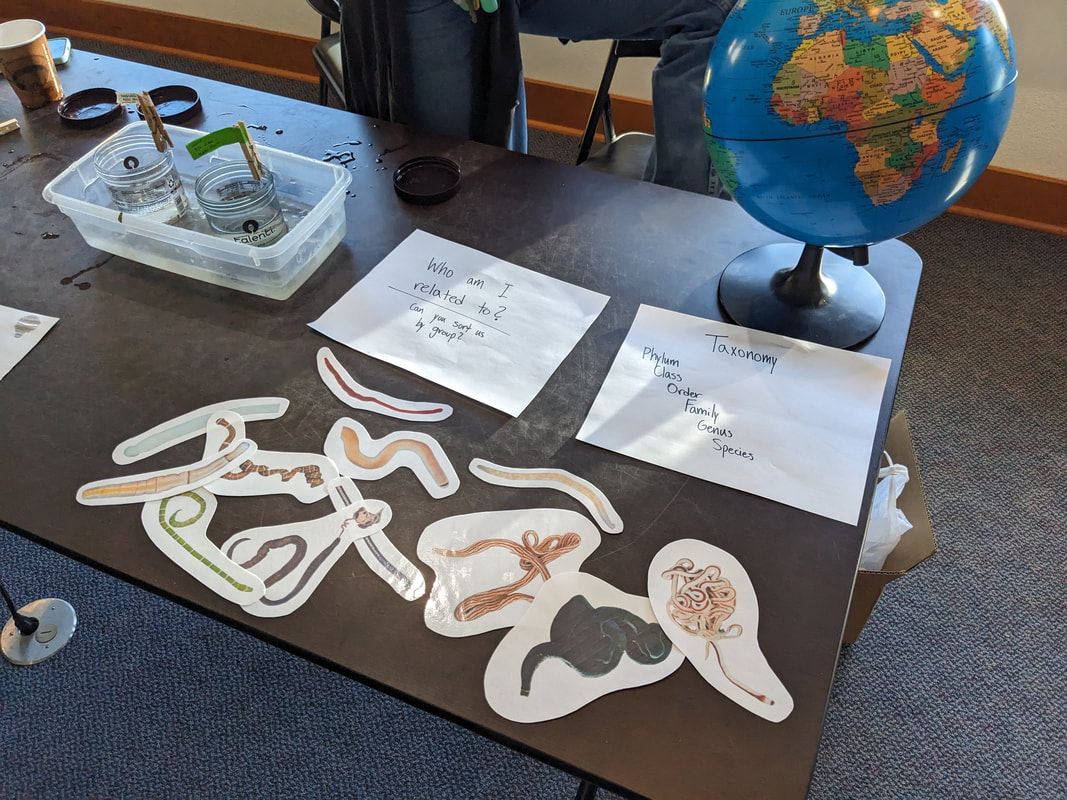
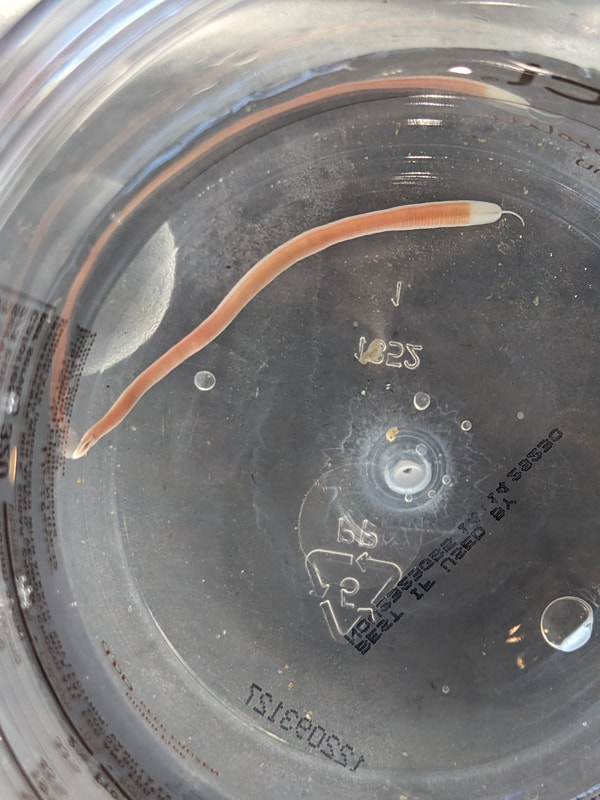
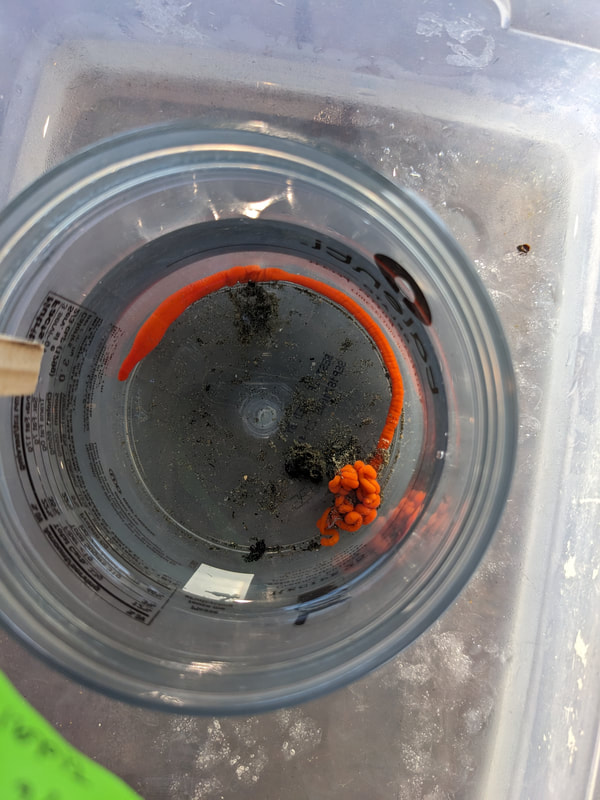
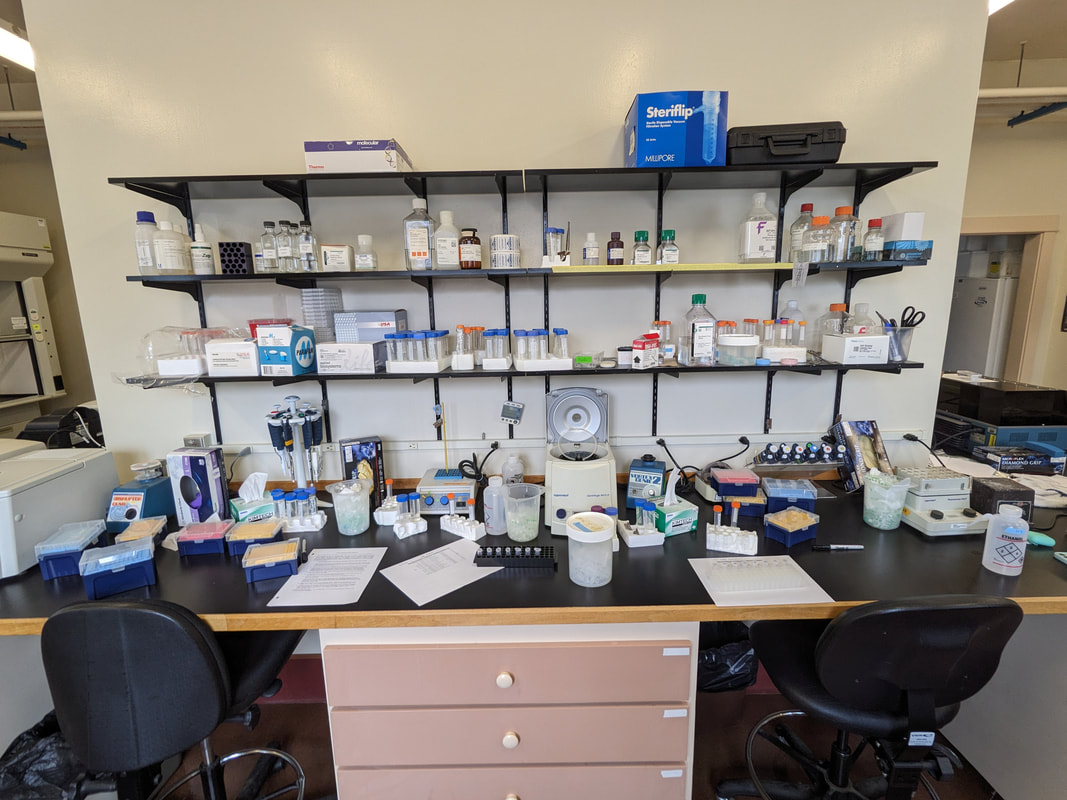
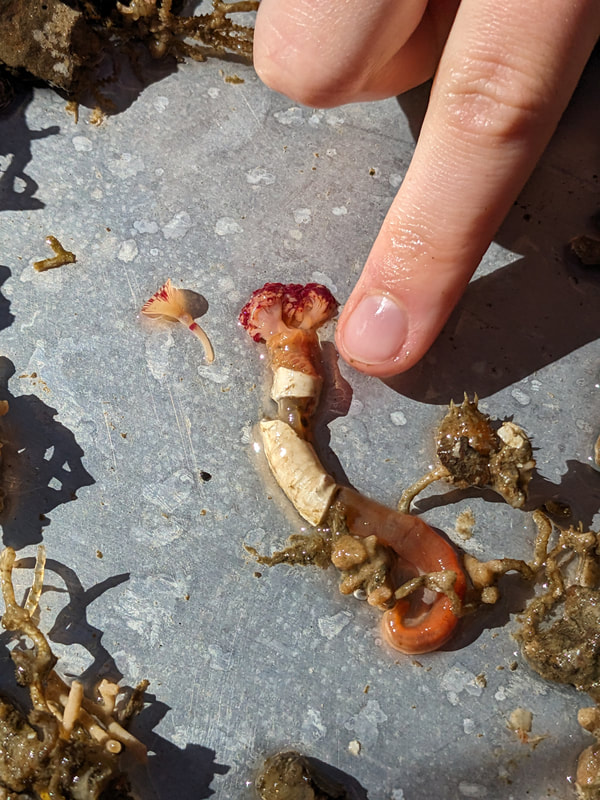
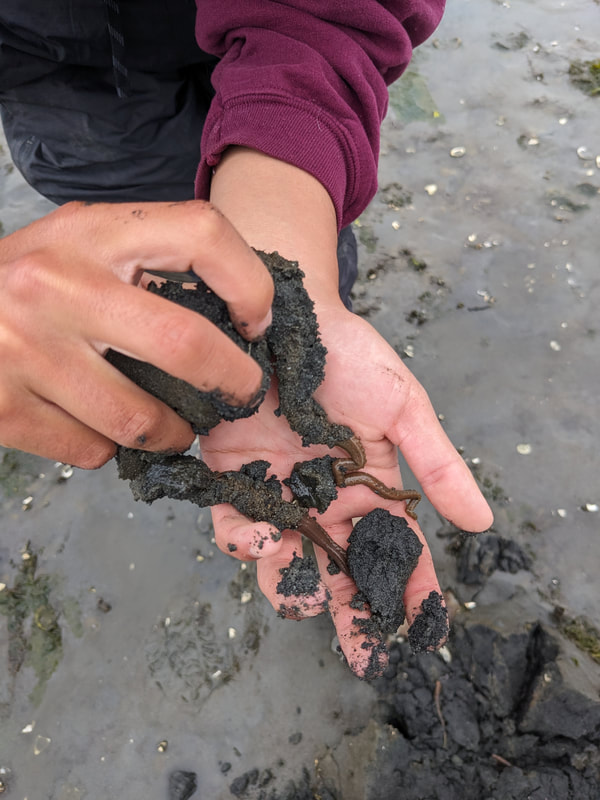
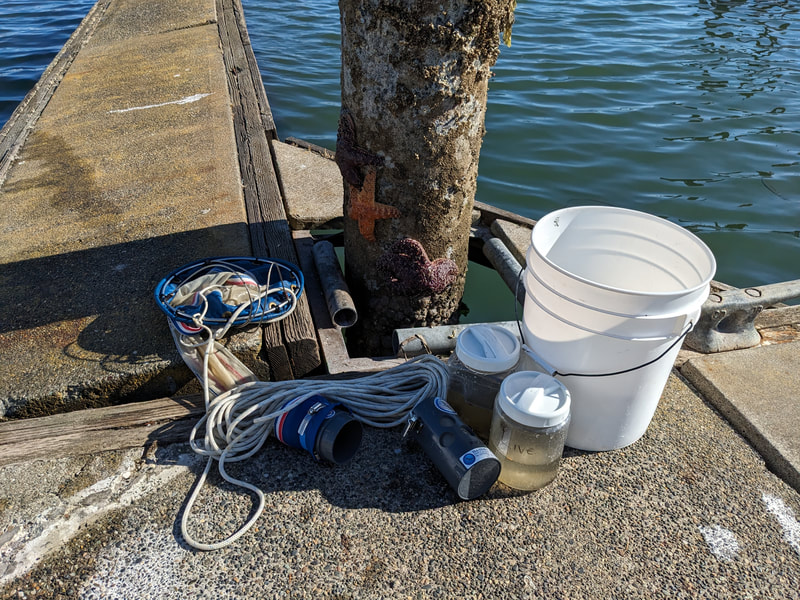
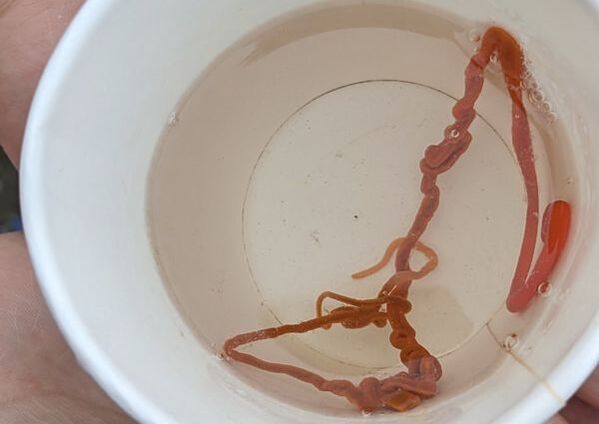
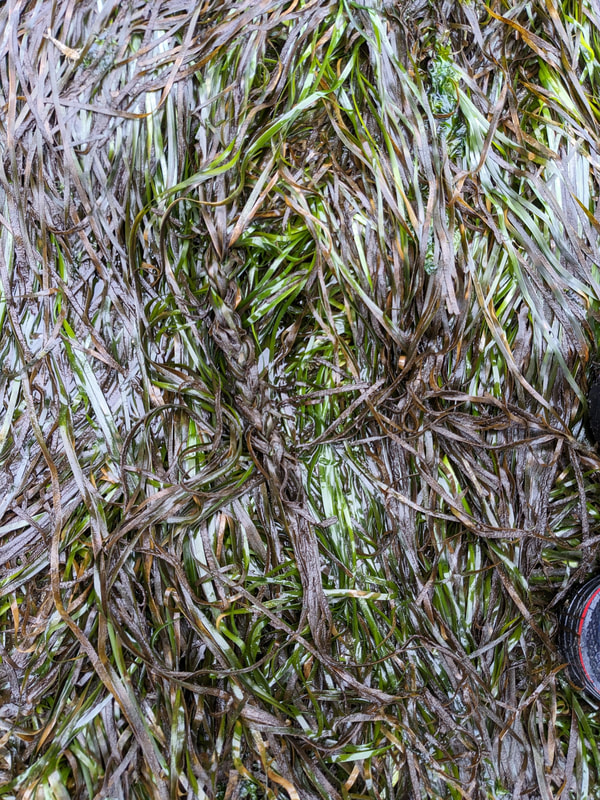
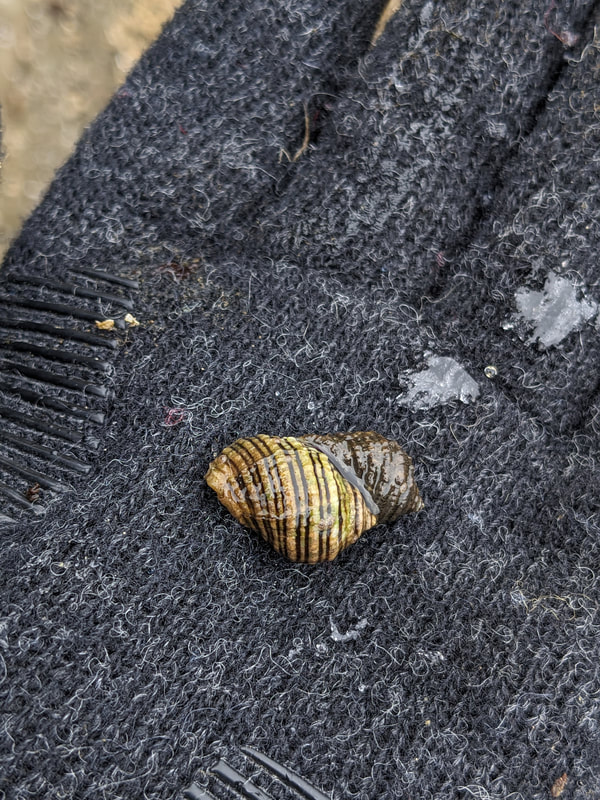
"I'm wandering in the mist just now. It'll blow off in a bit." -Margaret Renkl, Late Migrations Last weekend was an event I had been looking forward to, our public presentation at the Charleston Marine Life Center. All the REUs who are still here (Madison, Randi, and Tara are off having cool adventures) tabled for a couple of hours to talk to visitors about our research projects. Audrey and I assembled a tableau of live, locally collected nemerteans, an unintentionally devious image sorting game, and a blackberry DNA extraction station. I enjoyed trying to communicate what, precisely, I have spent the last two months doing, and I hope the kids we met enjoyed seeing the worms (and in a couple cases, touching them). The DNA extraction and micropipette seemed to be a big hit, although I do wonder how well we explained the purpose of the exercise. Alas, the rest of my week was not so enjoyable. I came down with COVID on Sunday and spent the next few days moping about and generally feeling crummy. As I write this on Thursday, my fever is gone and I'm feeling much better, but I didn't have a very productive week. Because most of our remaining work is database submissions and polishing posters, I was able to do some work remotely, and Audrey kindly did all the sequence clean up on the lab computer for our final plate. Despite the setbacks, I joined in the poster review and critique session remotely on Wednesday, and I got some helpful thoughts from Dr. Watts, Dr. Emlet, Riley, and my peers. Other than that, I've just been fiddling with R code to compile all of the specimen data and create nice-looking graphs for my poster. Besides my poster, the one silver lining to this week was that I finally took the time to figure out how ggplot controls all the visuals and aesthetics of graphs, and I am excited for future opportunities to develop illustrative plots. It's a little difficult to believe, but I am approaching my last week at OIMB this summer. Our final weekend will be spent touching up our posters, and then Audrey and I simply have to clean up the lab, organize our data sheets in a decipherable system for future reference, give a nice poster talk, and they'll kick us out! I've never had such an intense, focused, and joyful summer, not to mention the gorgeous, sprawling land and sea that surround me, and I already know that I will miss this place, a restful eddy before the current carries me to new challenges and adventures. I love it, for one week more. "I do not know what I may appear to the world, but to myself I seem to have been only like a boy playing on the seashore, and diverting myself in now and then finding a smoother pebble or a prettier shell than ordinary, whilst the great ocean of truth lay all undiscovered before me." -Isaac Newton This week marks the end of our lab work this summer. We re-amplified samples which previously gave us low quality reads (which was, in a pleasant surprise, quite successful), and measured the concentration with fresh Qubit fluorometer standards. This machine is fairly straightforward to use -- DNA absorbs light at a known frequency, and by measuring how much light is absorbed, the fluorometer can measure even very low concentrations of nucleic acids or proteins (our assay tubes ranged between 0.0100 and 0.7000 ng/μL). With these precise measurements, Audrey and I were able to dilute each sample accordingly, load our final 96-well plate, and send it to the OSU Core Lab for sequencing. We must tie up our project, perhaps sooner than we would have chosen, but we have hammered an enormous dent of knowledge into the previously pristine ignorance of nemertean diversity around the Arabian Peninsula, to the tune of 33 novel OTUs and counting. In order to make this information accessible and useful for future researchers, collectors, and naturalists, we have been uploading our data to BOLDSystems, a database meant for DNA-barcodes from all life on earth. One interesting aspect is that BOLDSystems has its own algorithm to delimit OTUs, which is less conservative than our ASAP analysis, so it records a few extra species. This is an important reminder that DNA-barcoding of the COI gene is a good proxy for differentiating species, but it is not the determining factor. The biological species concept is based on whether individuals and populations can produce viable offspring, and populations which do not interbreed will diverge genetically, but there are few sharp lines in the sprawling, patterned chaos of biology. Also outside of the lab, Audrey and I have been trying for the past week to collect nemerteans four our upcoming presentation at the Charleston Marine Life Center. We went to the Barview mud flats with Randi, but we had little luck. After an hour of sieving, we found one medium Cerebratulus, which we broke, and a few tiny, indeterminate worms. Most ended up being annelids (the bane of my nemertean-hunting existence!), though one piece turned out to be a broken-off nemertean midriff, probably from Maculaura alaskensis. It was fascinating under the microscope, with its branching brown intestines, and squeezed in between each branch are ovaries, bulging with eggs, each dotted with a nucleus. The Cerebratulus broke quite catastrophically, and its proboscis squirmed out the gaping hole in its body, but nemerteans have extraordinary regenerative abilities, and we hoped it would survive. This was not to be, for we found it rotting a few days later and dumped it down the sea table drain. A trip the next day to Sunset Bay brought us the classic Tubulanus ruber, but little else. On Dr. Maslakova's advice, we went once more to the mud flats, seeking small, brown, filamentous nemerteans that slink back to their holes as the tide recedes. We found one... and it broke and soon died. I now understand all the complaints I read from the 1800s about worms fragmenting before they could be properly studied. However, we went out on the search one more time today, this time with Dr. Maslakova. We chiseled barnacles off the dock pylons, and discovered quite a few Emplectonema viride. We are very pleased with these specimens, and will bring them to our presentation. I am also grateful for the opportunity to learn about collecting nemerteans, because there aren't many other people in the world who could show me so much about the practice. This week was also the annual Invertebrate Ball, which was a blast! We dressed up as cool invertebrates, and there was a swarm of impressive costumes. I am the proud winner of the Best Invertebrate Walk/Crawl/Strut/Move award. With lab work finished, these last two weeks will be a whirlwind of producing a poster, identification guide, and database submissions, as well as cleaning up our data, spreadsheets, and DNA samples to leave everything in neat order for future documentation. I'm both nervous and excited! Certain Knowledge from Herodotus These are the facts about the fish in the Nile: My favorite part of the past week was interacting with the South Slough National Estuarine Research Reserve. I went with Naia, Audrey, and Taylor to run an informational booth at the Coos Bay Farmers Market. We discussed the invasive five-spine crab (green crab), as well as the reserve's various programs and restoration work. I also enjoyed exploring the market, where I bought some nice plums and met Side of the Tide, the South Coast Folk Society's Morris dancing troupe, who seemed to be having a great time. Thursday morning, I woke up early to help South Slough collect eelgrass seeds. We tromped over to the Barview mudflat, which is just across the slough from OIMB. It is not heavily trafficked, but it was a beautiful and productive site. My duty was to extract mud cores along two 100 m transects and plop them into Ziplocs, and then sieve out the organic material from each one. I was very grateful for the borrowed wetsuit which kept me clean and (mostly) dry. I also saw several of the Morris dancers kayaking past, which was a pleasant surprise.
On Wednesday, we received the sequences from the two plates that gave us so much trouble last week. Our rate of high quality sequences was lower than before, but three-quarters were still good enough to process, which hopefully foretells more species discoveries. We received new standards for our Qubit fluorometer, although we haven't tested them yet, which we hope will allow us to measure the concentration in samples which did not sequence well.
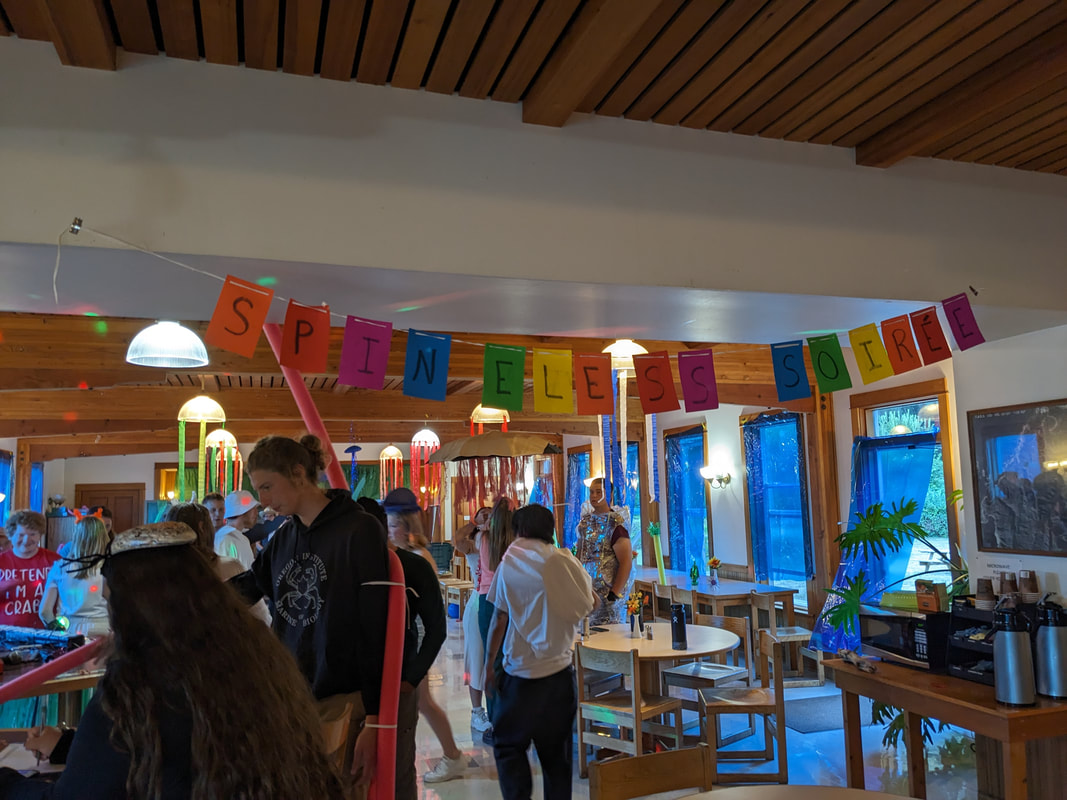
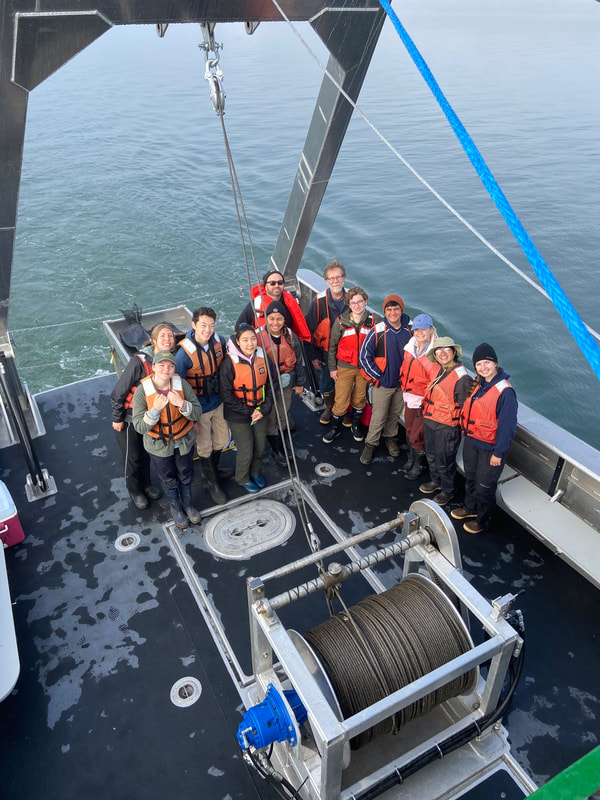
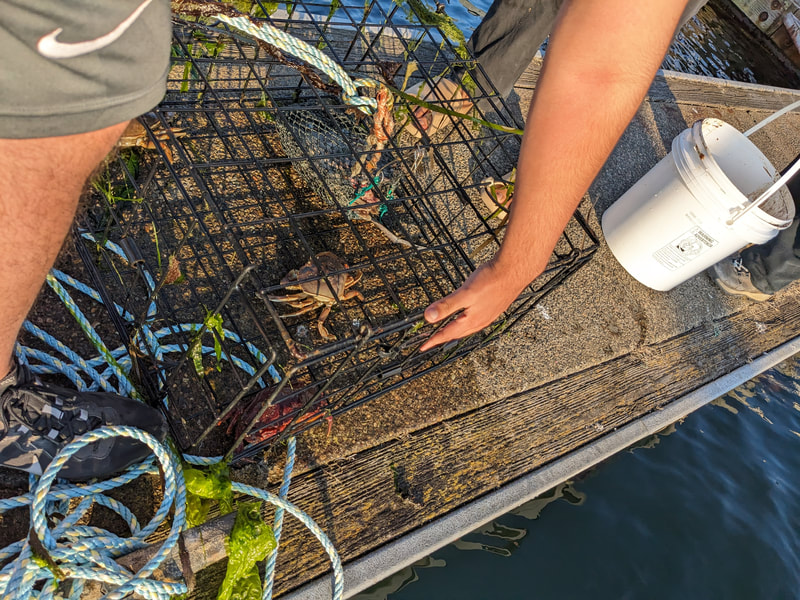
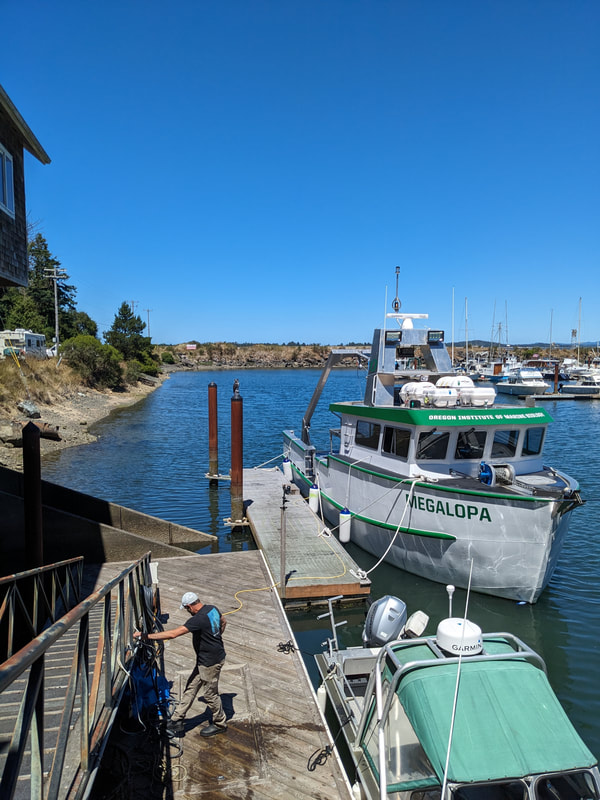
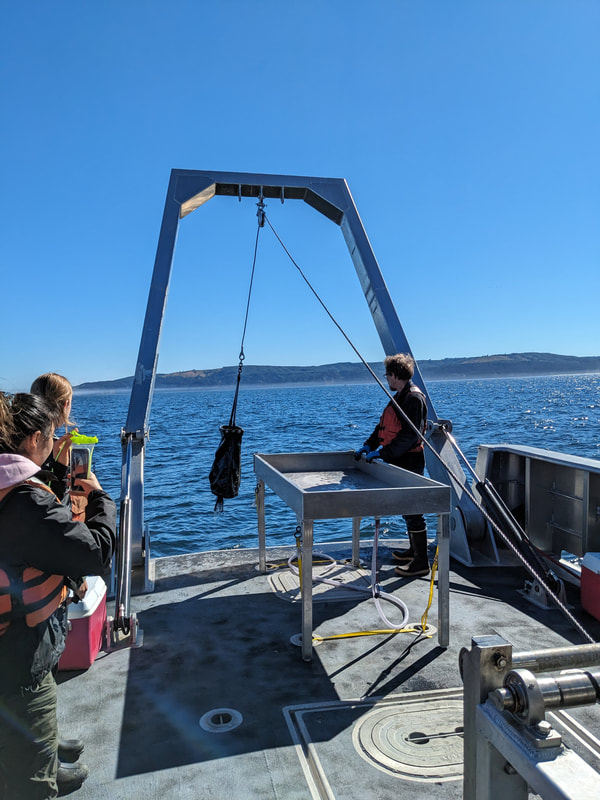
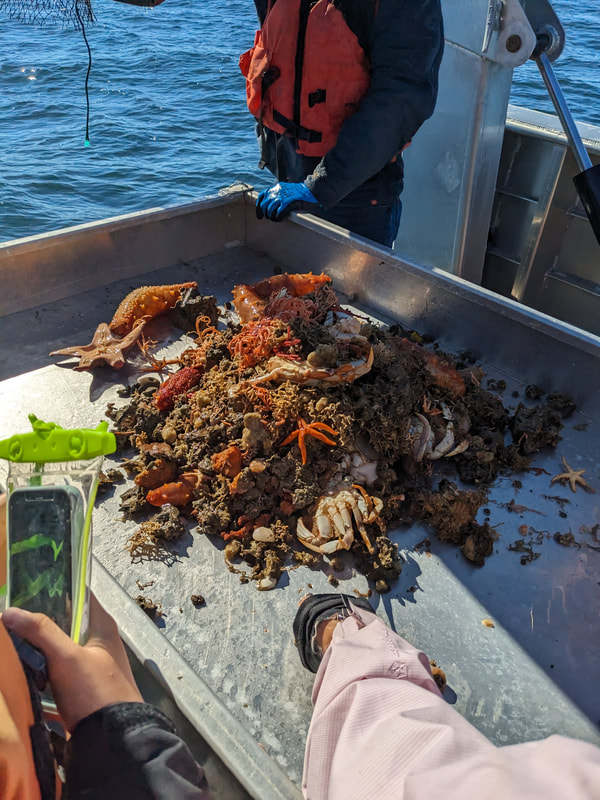
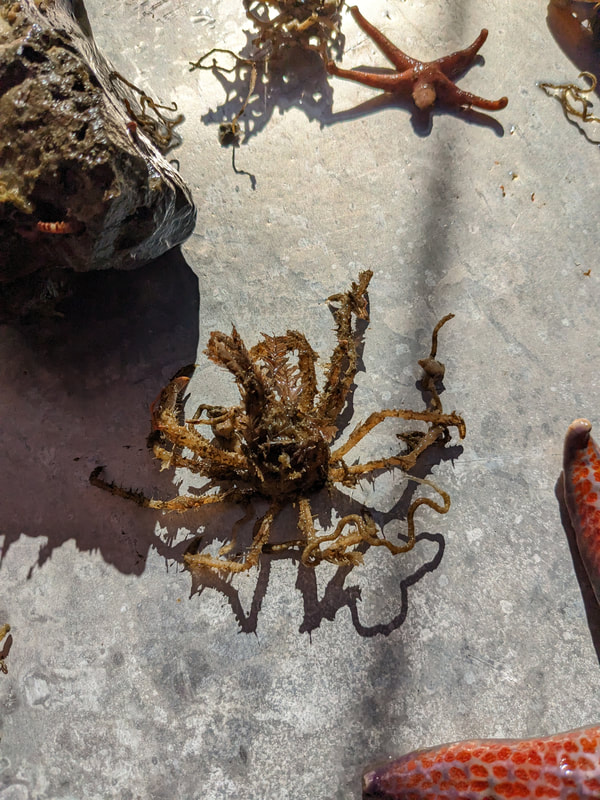
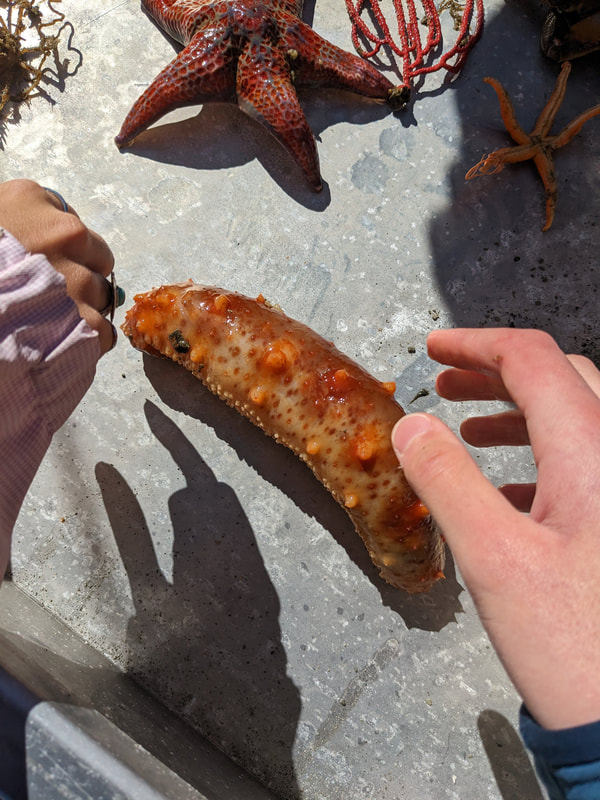
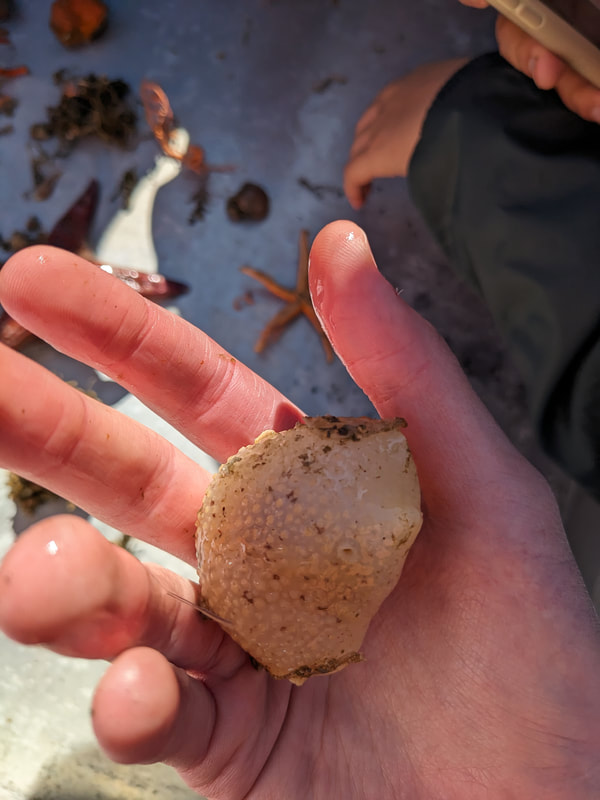
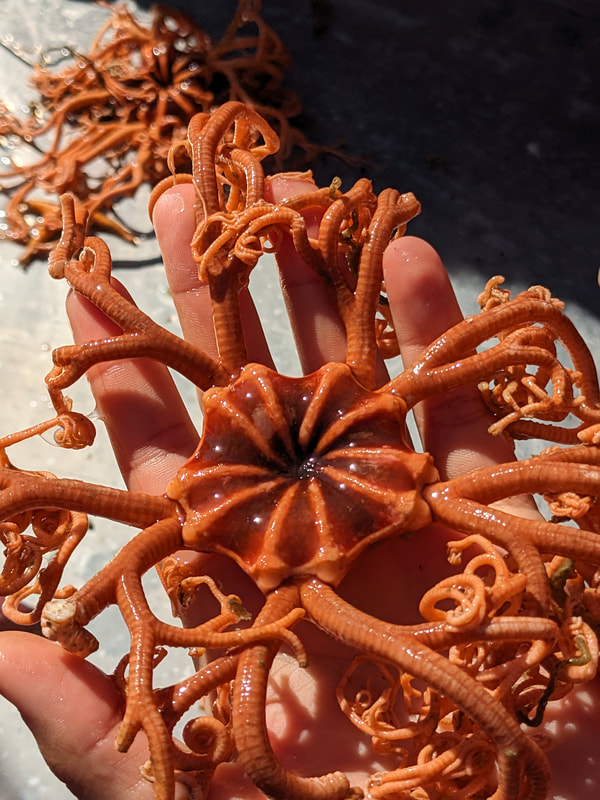
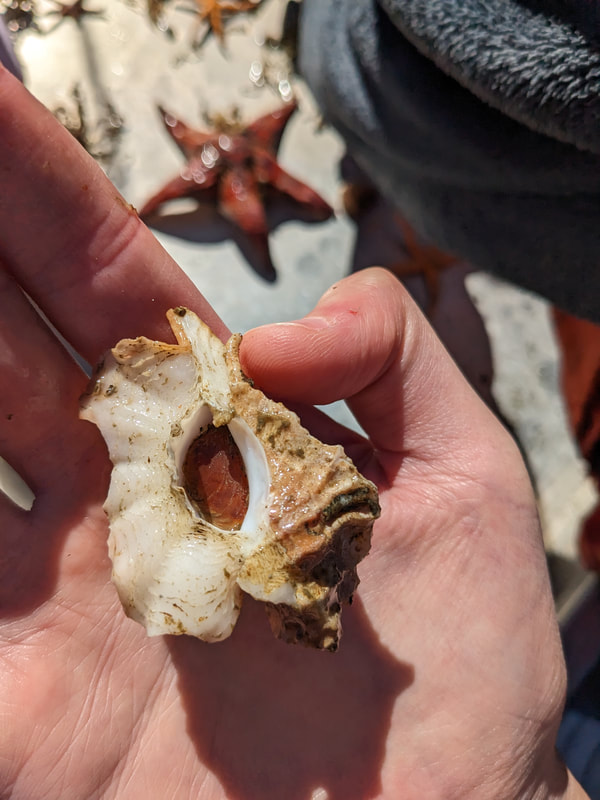
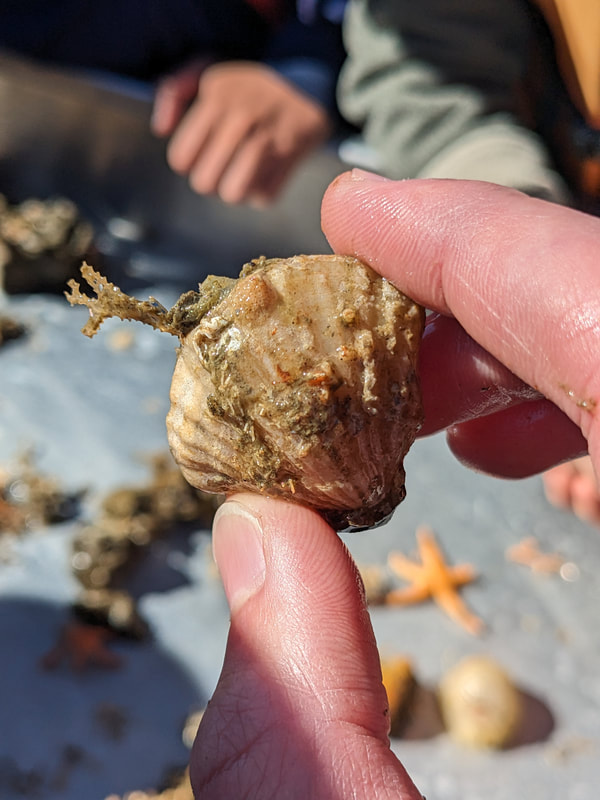
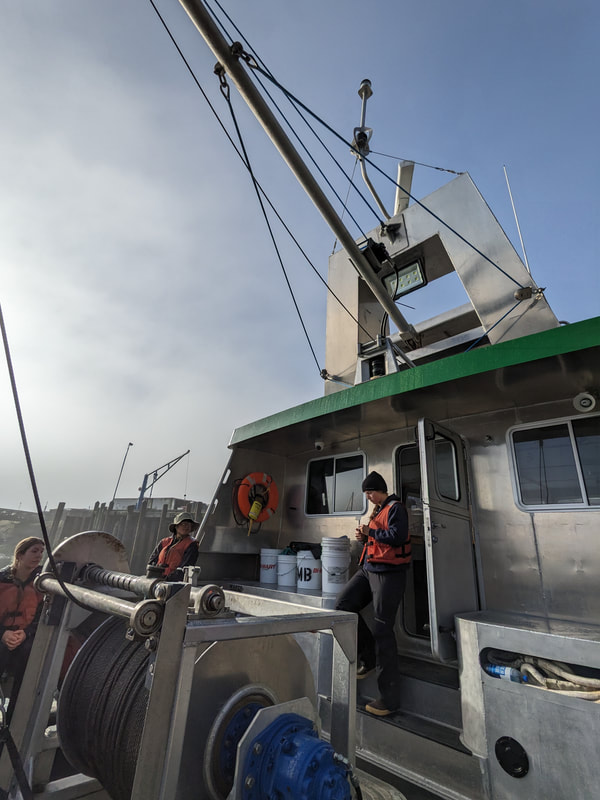
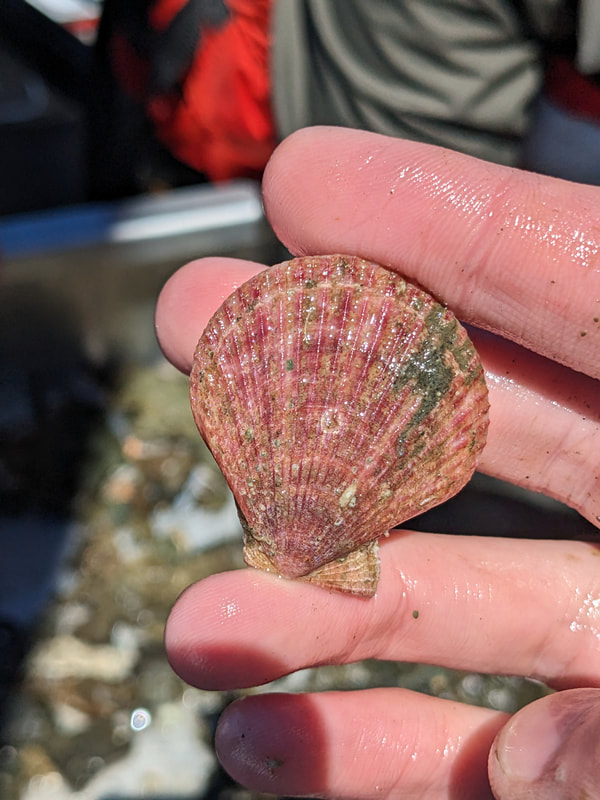
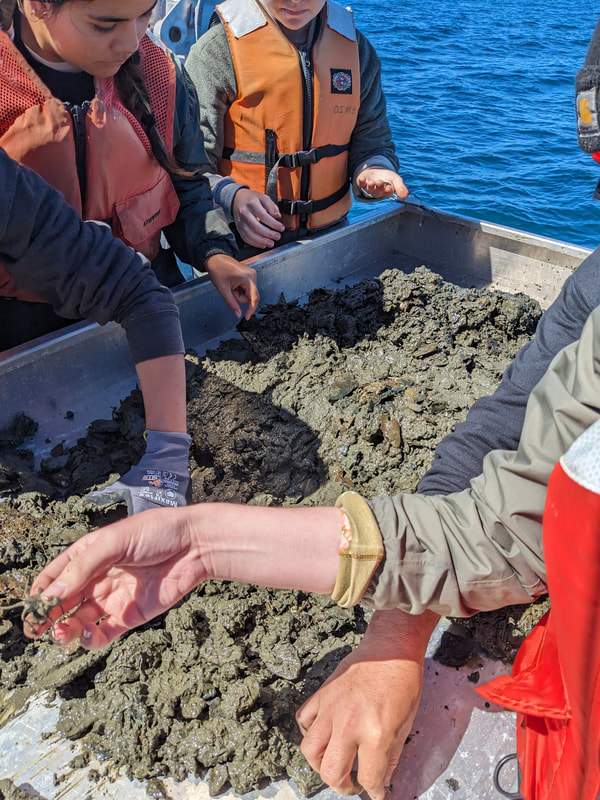
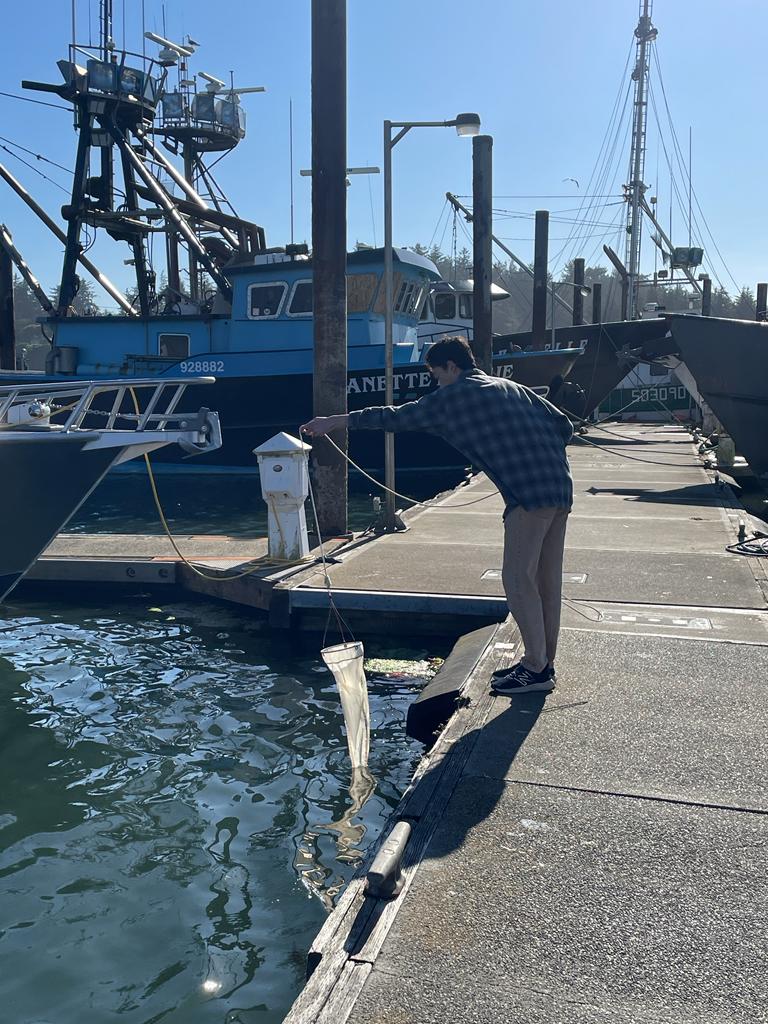
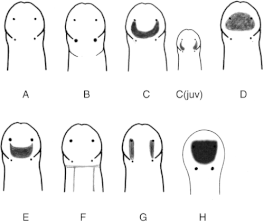
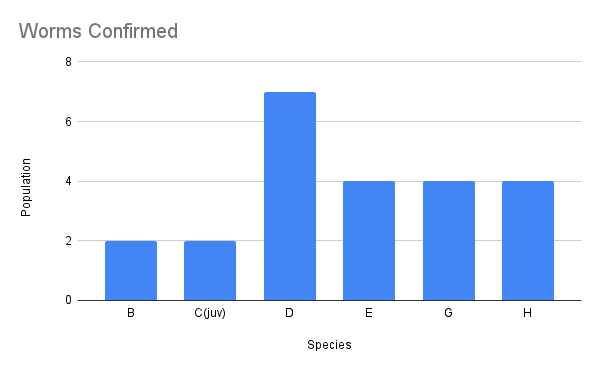
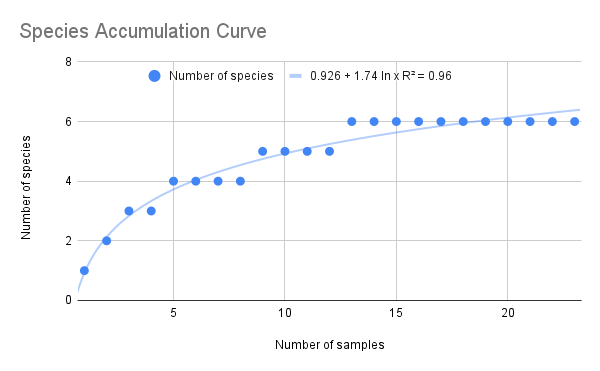
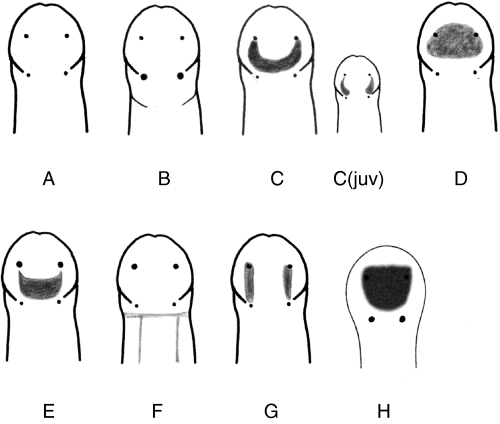
We have also been submitting sequences and associated specimen information to BOLDSystems, but have encountered a few processing issues. The biggest contention is that they would like to simplify our taxonomy, by moving information about which operational taxonomic unit a specimen sorts to or which species it resembles to the notes column, instead of the species column. We'd rather keep this information prominent, because it is important to have a unique identifier for every operational taxonomic unit. The open taxonomy notation ("this specimen is similar to such-and-such species, that one looks like that species") is necessary, since so many nemerteans are undescribed, and we have no other way to anchor unique identities. As we head towards Week 7 of 9, I am feeling the time crunch -- I want to collect all possible data from every sample, submit information to databases, create detailed ID guides, and prepare engaging presentations for the general public and for our poster session, not to mention explore the wonderful Oregon coast, but our time here was always going to be limited. I hope I will find a satisfying conclusion to my project and appreciate the last few weeks for all they are worth. Or, A diſcourſe on the abſence of monotony, being a time of inopportune distreſs, initiated by forces outſide the author's demeſne "Who are you, who are so wise in the ways of science?" -Sir Bedivere, Monty Python and the Holy Grail Saturday was the coolest day -- we went dredging on OIMB's shiny new research vessel, the R/V Megalopa. Dr. Emlet was an ace at identifying the many organisms we pulled up from 150 to 300 ft deep, and the incredible crew (Captain Bradd Beckett and James Johnson) made the trip smooth and enjoyable. The boat is equipped with an A-frame to support the small trawl net, and I operated the winch for a bit. Several people got a bit queasy, including me, but no one lost their lunch, since it was a fairly calm day. What helped me most were ginger chews, saltines, and sitting in the cold wind, so I'll come prepared in the future! The basket stars we pulled up were magnificent, but our table was overflowing with over a dozen phyla: sea cucumbers and brittle stars, corals, a Cerebratulus nemertean, sponges, jellyfish, sipunculids, fish, Dungeness crabs, nudibranchs and chitons, serpulid tubeworms, phoronids, bryozoans, brachiopods, scallops, and several more I must have forgotten. We presented about our projects to the other REU students on Wednesday, and it was really cool to see what everyone's been up to! It was revealing that everyone thinks their own project is monotonous, but everyone else sees how cool the research is. "Thus took form by degrees those hundred thousand diverse races, the classification of which so cruelly embarrasses the unfortunate race that habit has changed into naturalists." -Georges Cuvier, Recherches sur les ossemens fossiles de quadrupèdes As for my project, it has been anything but monotonous. It's hugely exciting to see how many ways research can go wrong. Since last Friday, we have passed 670 amplification reactions, but there are still 24 samples (out of 180 that are new this summer) which we can't amplify. We have a few last tricks up our sleeve, but many of these will remain unsequenced. This is disappointing, but today Dr. Maslakova reassured us that there is always a percentage that cannot be sequenced, and moreover, with or without these samples, our project is groundbreaking in documenting nemerteans from the Arabian Peninsula. Her outlook is both pragmatic and optimistic, for when we have made every reasonable attempt, we must be able to cut our losses and focus on the many other tasks and endeavors that are always begging for our attention. Banging your head against a wall over and over won't break the wall -- just your head. However, we were enormously excited this morning to have 96 samples loaded into two plates to send for sequencing -- when we discovered that the sequencing company the Maslakova lab has been using for years permanently shut down last week. This caused a real scramble. The University of Oregon Core Facility doesn't offer Sanger sequencing service, but the Oregon State University Core Facility doesn't measure and adjust the DNA concentration, which is necessary for a high quality read. In the past, the sequencing company would quantify samples for us, so we weren't prepared for the procedure. We tried a Qubit fluorometer, but the standards wouldn't read properly. We also tried the gel-quantification method, which uses a ladder with DNA fragments of known lengths and concentrations. The ladder we borrowed from Dr. Emlet did not give a very clear image, but it was enough to roughly determine the concentration of test samples, by guesstimating how much brighter or dimmer each sample seemed compared to the ladder. We now have to go through our gel images, decide how bright each successful band was compared to our test samples, remove our products from the loaded plates, dilute them by a factor between 1 and 12, and load them into a different layout, plus primers. This promises to be a time-consuming and monotonous process. Alas, we have no other option, and we desperately want to get these 96 specimens sequenced, so we're going to go through with it. I am experiencing firsthand that science can be tortuous and arduous, but with luck, we will send our samples for sequencing next week, and soon crest back atop the emotional high of species discovery! "I must have been distracted when I left my home because P.S. I wrote this blog just after our dies horribilis, but since Friday we have finished loading up the new plates. It was a lot of work, but we are prepared to send them for sequencing on Monday, and I am very excited to see what we will discover next week! ...which must in time find the sea I look back and can't believe it's already been a month here. We submitted our research proposals to the faculty organizers of the REU program, Drs. Maya Watts and Richard Emlet, for feedback this week. Next week we'll present our projects to the other REU students, so we're really in the thick of it. Also great news, I have finally met Dr. Emlet in person! He has returned to OIMB after teaching a course at Friday Harbor Laboratories up in Washington, and at lunch today he had some very insightful advice about career paths and the state of academia. Last weekend we hit up a string of excellent thrift and vintage stores in Coos Bay, where I picked up a flannel and a book of Ansel Adams prints. That evening we went to the 7 Devils Pub and Brewery for a "Three for Silver" concert recommended by my mentor, Dr. Maslakova, and her husband, Dr. von Dassow. He knows the band particularly well, since he once hosted them as artists-in-residence for two weeks in his lab. They were real crowd pleasers, and I had a great time dancing by the fire. Audrey and I have been emerging from our lab occasionally to poke around in the inter- and subtidal. We joined Jay and Adam to count the daily catch of the Dungeness crab megalopae (spiky larval forms), and it was fun to see the long-term monitoring that can happen at a marine station. We also collected a plankton tow with Chloe last Tuesday, which is a simple but useful technique. Later in the week, Chloe came into our lab for a little research exchange, pipetting a ladder in our agarose gel. Thursday morning, we went with Randi, Madison, and Chloe on the mud flats to muck around with the phoronid worms, clams, and polychaetes. As I write this, we just got back from a new hike with El -- literally a new trail that a couple students hacked up the coast. It's still rough in spots, but the mist was beautiful today, and I love investigating these small pockets of wilderness. Audrey and I are, however, still doing plenty of lab work. We received our second plate of sequences, and after hours of cleaning data and sorting spreadsheets, we have a grand total of seventeen tentative species newly sequenced this summer (and likely new to science!). This is a pretty incredible haul, and we still have at least one more plate of samples. We compare each sequence against the several hundred samples Dr. Maslakova has collected and the GenBank database. Assuming the attached identifications are correct (which is not always safe to do), the matches suggest what taxon we have sampled. The worst mismatches to decipher are the subtle ones, which sort to the wrong genus or clump visually disparate specimens together. This could be because of mislabeling in the lab or very surprising phylogenetic similarities. Assuming our DNA extracts are uncontaminated, we can re-run amplification to get more accurate results. However, some of these samples are likely irretrievable, which is always a risk. We'll keep chugging along, testing different tips and tricks, and I hope I'll have good news about them next week. I know everyone is eager to learn the worm poll results. This highly scientific survey shows that one third of my audience is Tetrastemma melanocephalum (shadow face). There are also quite a few T. vermiculus (crying) and T. longissimum (aaahhh). Also represented are T. peltatum (smiley face), T. laminariae (choker with large dots), and juvenile T. coronatum (sad baby). Species richness: 6. Average population size: 3.8 ± 1.8. Simpson's Inverse D: 6.17. The species discovery curve shows that I can expect to discover about 10 species of Tetrastemma in my audience if I sample 184 respondents.
No matter what I say,
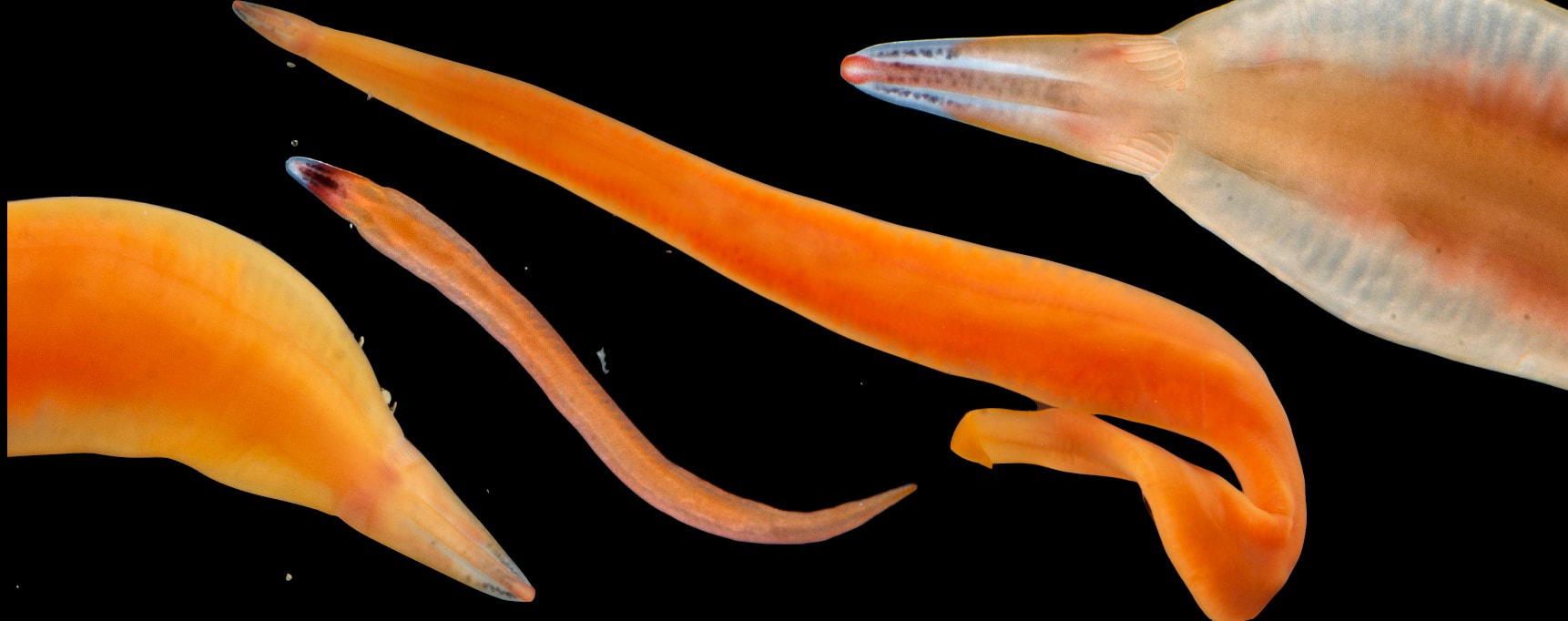
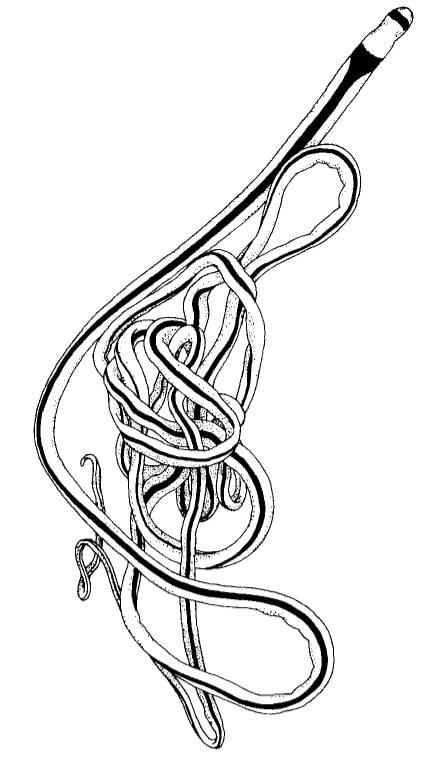
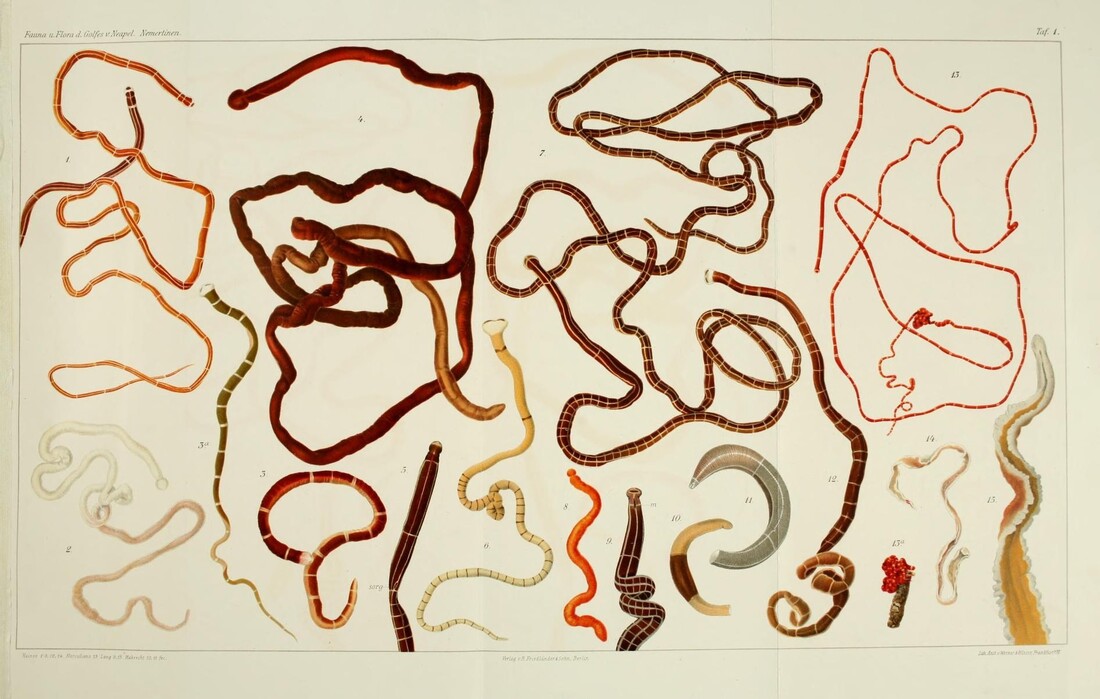
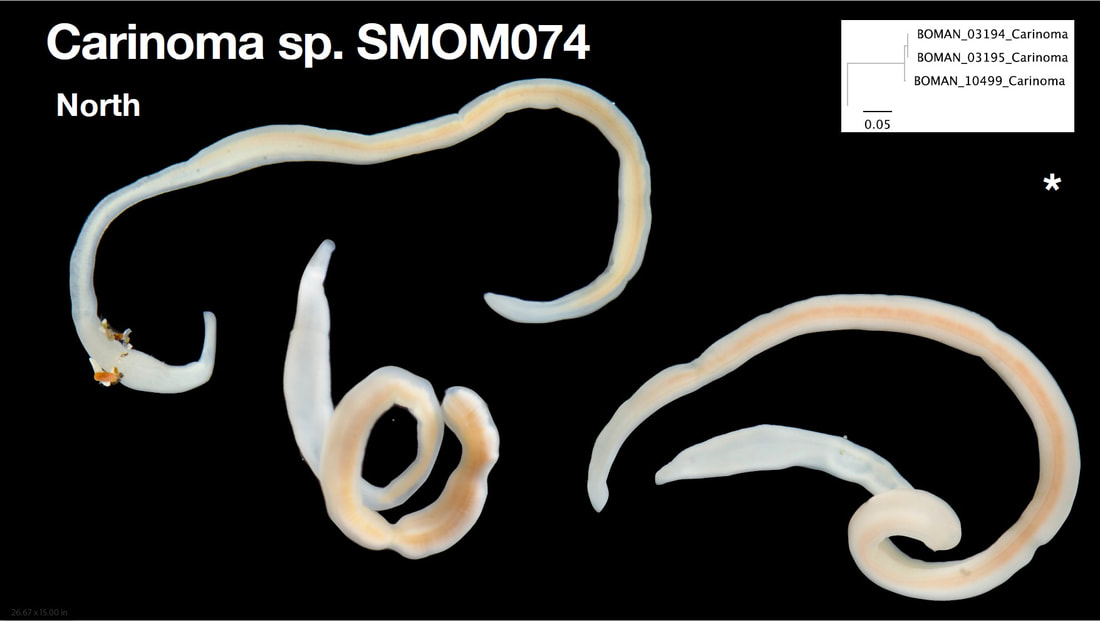
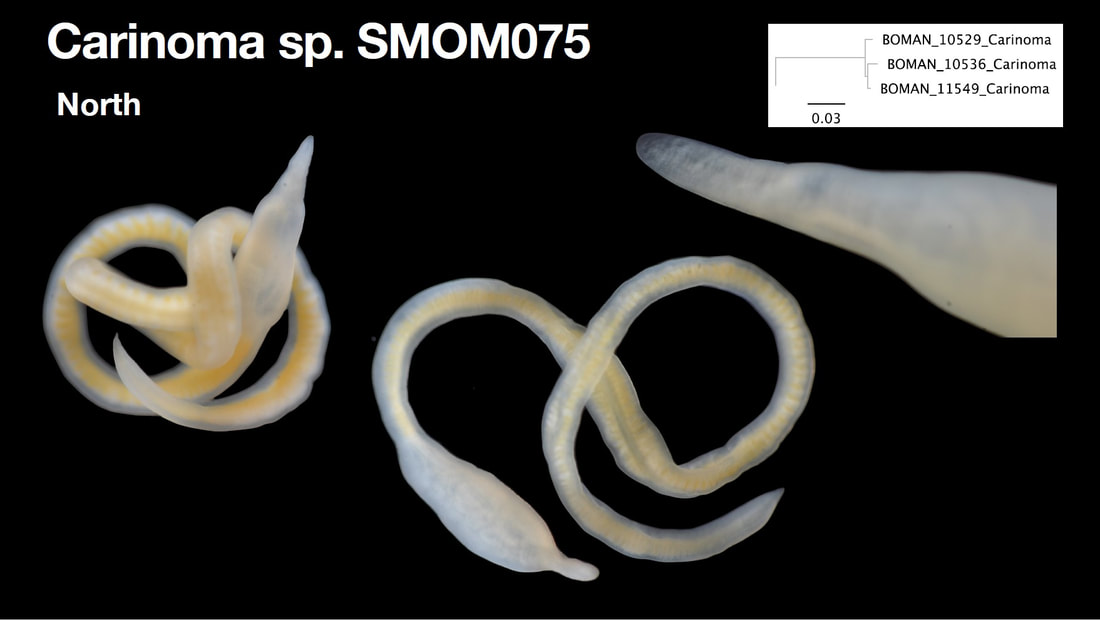
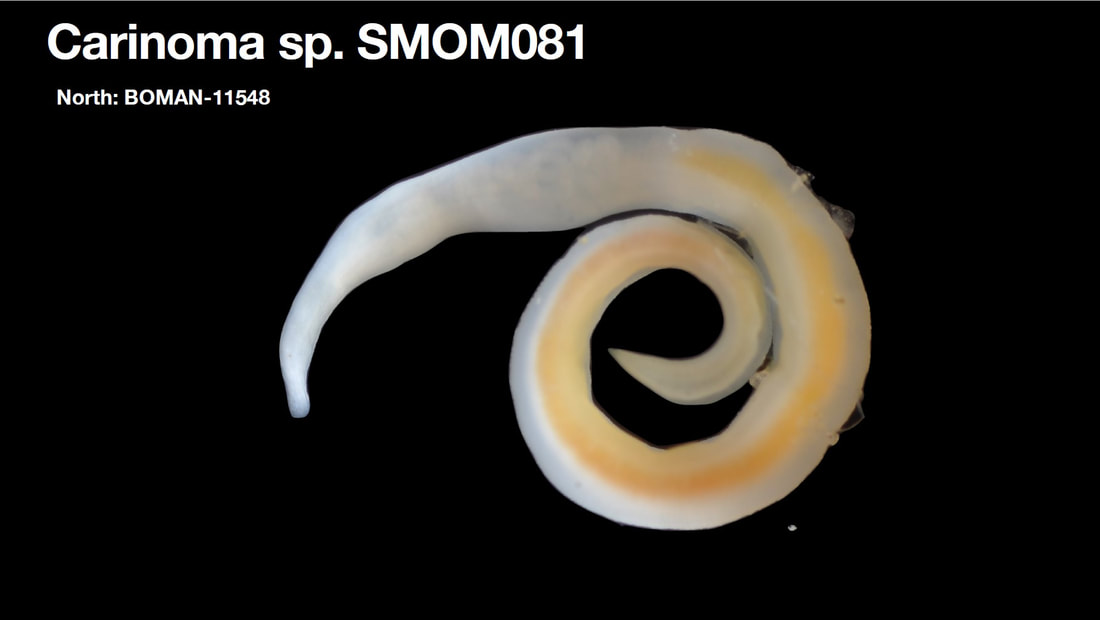
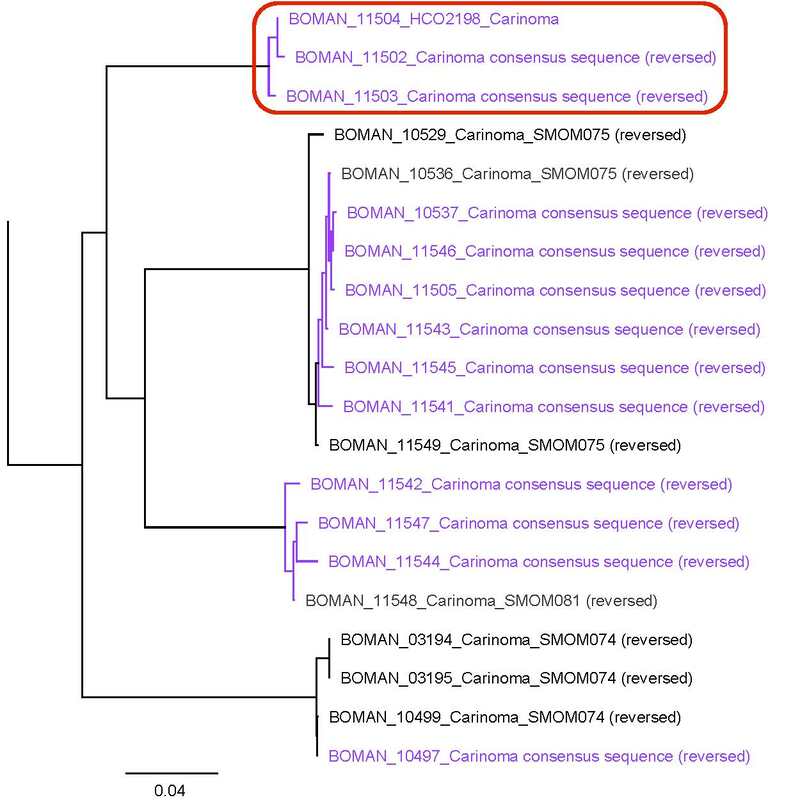
Saturday night, we gathered around the fire, ate some innovative cookie-s'mores proselytized by Chloe and Audrey, and tried to tell ghost stories. El opened with a fabulously spine-chilling campfire tale, but no one else knew any good ones, so I told a very silly joke about a village of elves, an old wise wizard, and the frightful Medikin. If you're curious and have 15 minutes to spare, I'll tell it to you someday... I'm working on my research proposal, which includes seeking out original species descriptions, and there are some crazy illustrations! These are serious business. Wormy squirm. Lab work this week has been super exciting, because we received our first 48 sequences on Monday! With a nifty program called Geneious Prime, we first trimmed off the ends where we saw low quality reads or primers, which don't reflect the nemertean's actual genetic sequence. Since we sequenced from forward and reverse primers (one on each DNA strand), we had two reads of each region, which Geneious Prime aligned and searched for discrepancies. We then BLASTed the sequence (a great opportunity for fun sound effects *whoosh* *blaaaast*), meaning a search against a national database called GenBank for similar genetic sequences. After checking for mismatches with initial field identifications, which would suggest contamination or mislabeling, we constructed a phylogenetic tree to see how our sequences grouped, and it was awesome! Some species overlapped with previous specimens, which was both expected and good confirmation, but we also found 5 entirely new, never-before-sequenced “operational taxonomic units” (OTUs) in just the past three weeks! These OTUs are genetically distinct enough that we believe they are reproductively isolated populations, a.k.a. separate species. Here's a sneak peak at our results, just using the couple that I have images for: One specimen groups genetically with an OTU identified to the suborder Reptantia, and visually they are also quite similar. On the other hand, this one represents an entirely new species of Cratenemertidae. Here is another specimen matching a Lineidae OTU. But it's important to remember that a big reason we are using genetic techniques is because of cryptic species. Here are three OTUs within the genus Carinoma that are visually indistinguishable but genetically distinct. Here's the branch of our phylogenetic tree for Carinoma. Purple sequences are ones Audrey and I just added. Most of them group closely with one of the three OTUs documented from Oman (pictures above), but the ones boxed in red are completely separated -- a new OTU! I don't have pictures for it yet, but those worms probably look pretty similar to the other three tentative species. So hopefully this gives you a better idea of what we're trying to identify, why we're doing it, and what we're discovering!
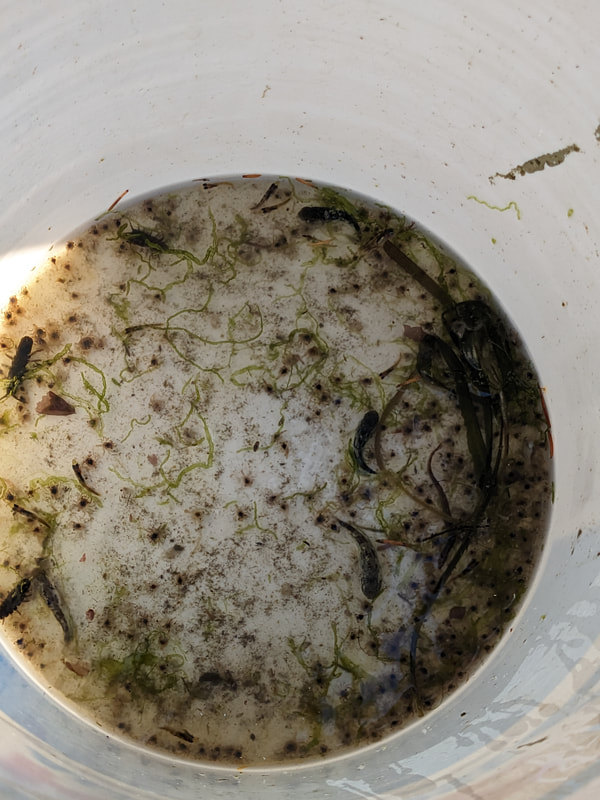
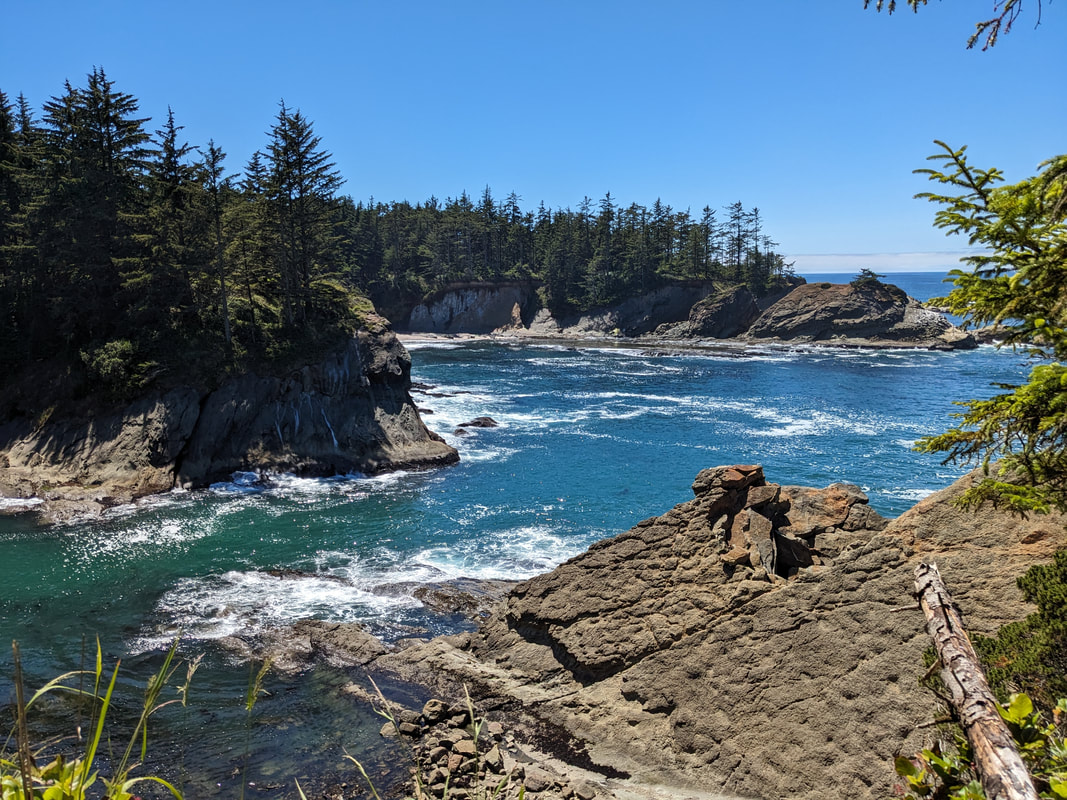
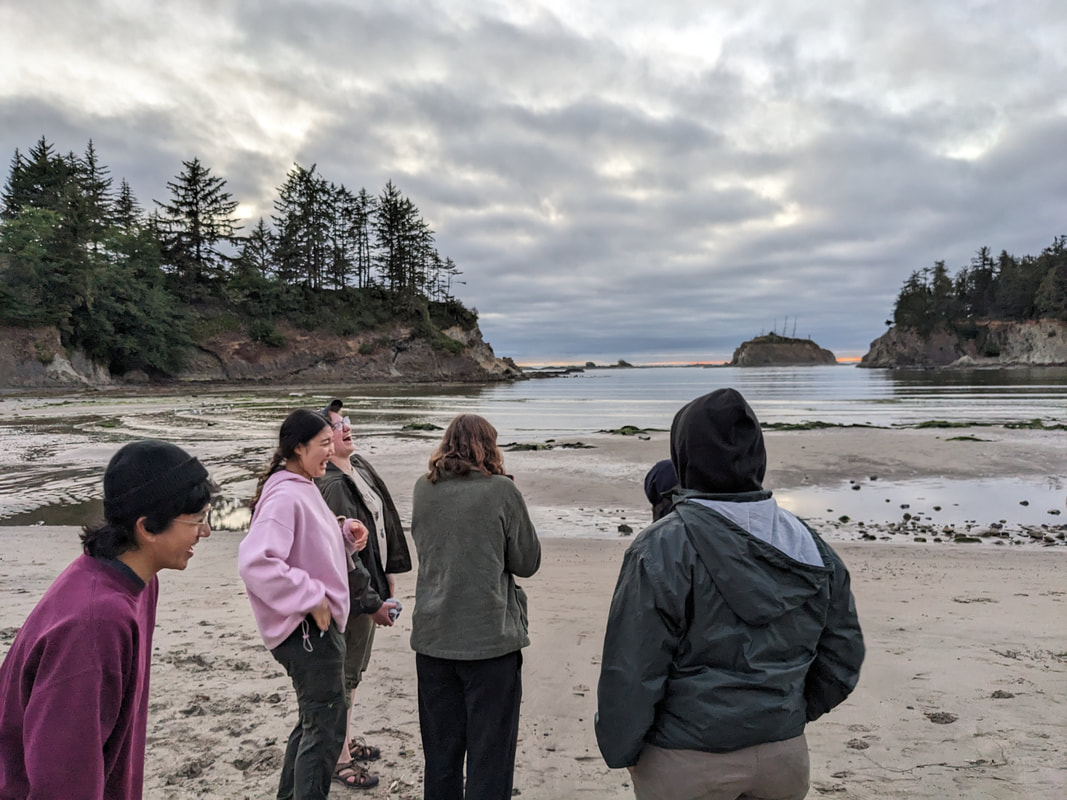
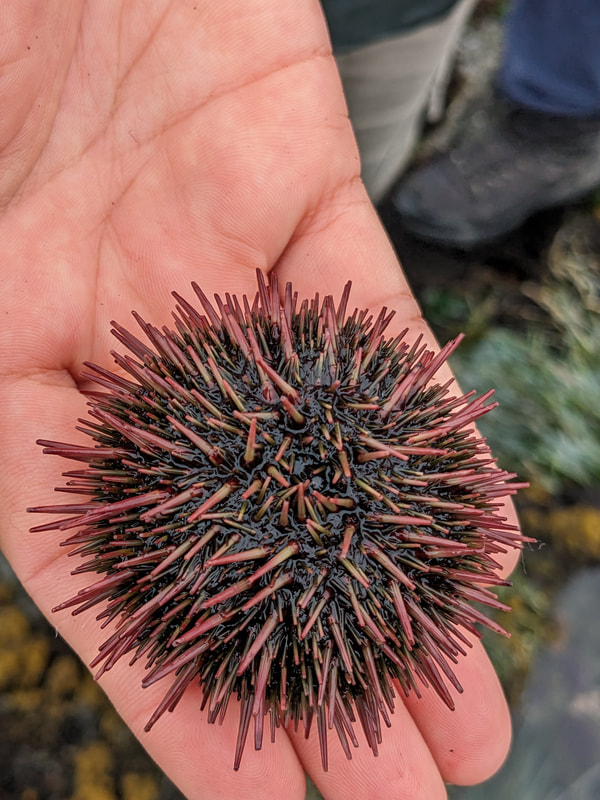
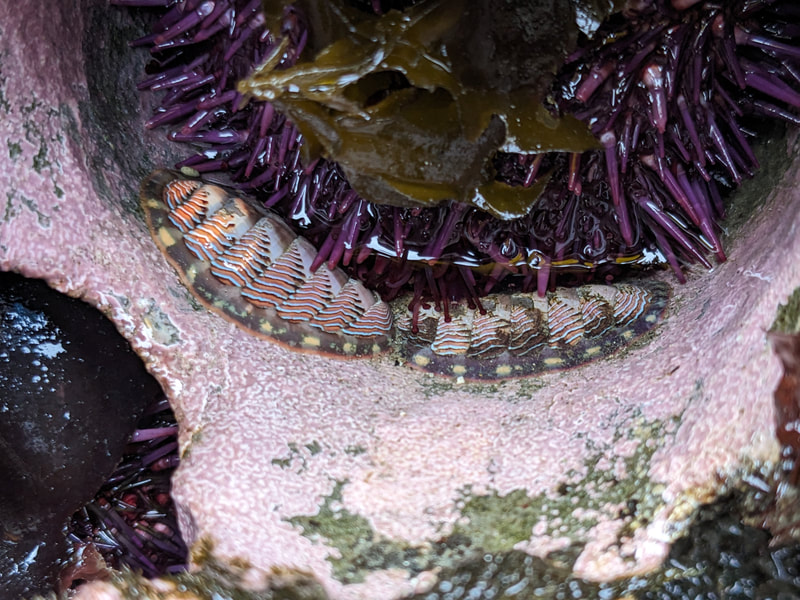
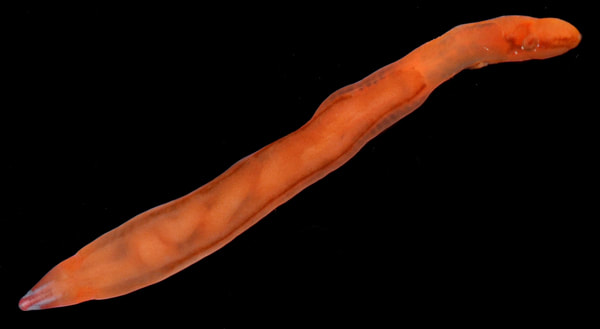
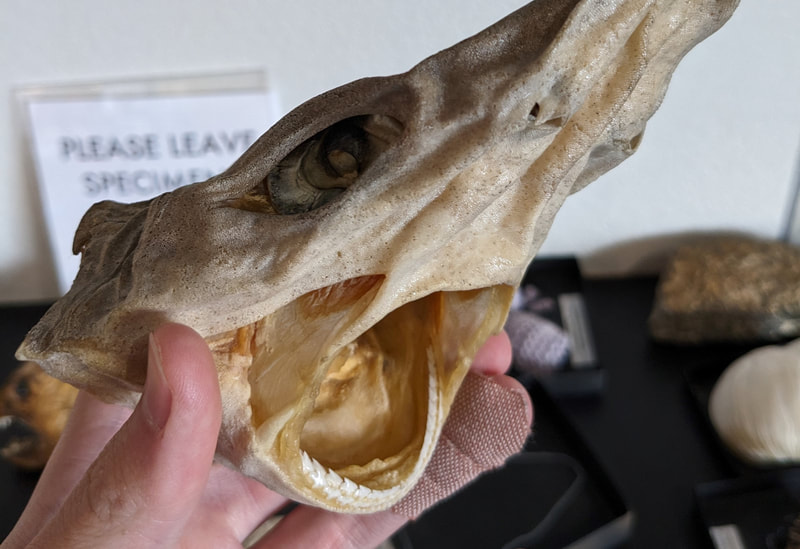
You can really have no notion how delightful it will be My highlight this week was tidepooling during the spring tide, when the sun and moon align to create dramatic changes in water level, which gave us a strong -2.2 ft low tide. I went with Riley (our awesome REU coordinator), Randi, Naia, and El to Sunset Bay to poke around in the eelgrass beds and rocks. There was lots of scrambling, down the cliff and then across algae-covered rocks, sea urchins, and eelgrass. I slipped a couple of times, but luckily I got off with only a few scratches. Besides the abundant purple sea urchins and ochre sea stars, we spotted red sea urchins, gumboot and leather chitons, six-rayed sea stars, sea slaters and kelp isopods, some very pretty nudibranchs, and a bright orange Tubulanus ruber nemertean worm. I got pretty cold after a couple hours, but it was an incredible experience, and I'm very glad I went! Tuesday was the Fourth of July, so us REUs went to the city of Coos Bay to watch the fireworks. We were also planning to wander around and find something to eat, but almost everything was closed, so we ended up at McDonald's, which was oddly apropos. On the way back, we had a lot of trouble wrangling the van, first with the parking brake (who knew you needed to pull the release handle?), then with turning on the headlights, then with turning off the roof light I had accidentally turned on while trying to find the headlights, but we made it back okay. On the research side, we have finished the first pass on all of our nemertean genetic samples currently on-hand, which is great news! Unfortunately, we are now in the dreaded troubleshooting phase, with finicky samples that did not work initially, for several possible reasons. Some nemerteans had a delicious meal of polychaete worms right before they were collected, so we ended up with a sequence matching the wrong phylum. Others might have enzymes bound up in their DNA which inhibit replication. Earlier this week, we used a 10% dilution of the DNA, which counterintuitively can improve results by diluting these enzymes, and in fact that technique solved about half of our problematic samples. For a third group, we may need to test different primers because of genetic variation in the target gene. Primers are short DNA sequences that provide the starting site for the replication enzyme. We usually start with the so-called 'universal' Folmer primers, but we moved into testing nemertean-specific primers in various combinations, which might bind better to nemertean DNA and improve the signal. Next week, we'll continue testing combinations of primers, a 1% dilution, or lowering the temperature during polymerase chain reaction to encourage binding even when there are slight mismatches to the DNA. In an exciting development, we mailed our first plate of samples to an outside company for sequencing today, and we should have that data back by Monday, which will allow us to start identifying tentative species. I'm looking forward to our first results! And there came of a sudden creatures with backs as hard as anvils, bent of claw, walking aslant, squinting, scissor-mouthed, shell-skinned, bony-natured, flat-backed, gleaming-shouldered, bandy-legged, with tendons for hands, peering from their chests, eight-legged, twin-feelered, unwearying: those known as crabs. -Batrachomyomachia (The Battle of Frogs and Mice) My first week at OIMB has been a whirlwind! It's a wonderful experience to live and work on a campus with a small community where everyone is captivated by the marine world. My first day on campus, a big group of students went down to the beach to poke around, and everyone was exclaiming in delight at the ochre sea stars, picking up crabs, or staring intensely at the little tidepool sculpins. Everything I need is nearby, from the dining hall and 24-hour library to the OIMBeach and everyone's favorite convenience store, Davy Jones' Locker. I visited the Charleston Marine Life Center yesterday, where the collections of living and preserved organisms are fascinating! Highlights include the Humboldt squid and the dogfish. There are also a series of trails running up the mountain behind campus, and it's impossible to convey how stunning the Pacific Northwest conifer forest is. My research is off to a great start -- from samples of ribbon worm (aka nemertean) tissue preserved in ethanol, I have extracted DNA, run it through a polymerase chain reaction (a pivotal method to amplify the specific regions of DNA), checked product quality in an agarose gel, and purified the PCR products. Early next week, when we have a few more samples to fill a plate, we'll send them off to an outside company and get our first sequences! With these genetic sequences, I'm going to look for which specimens belong to the same species and which have enough divergence to be separate species.
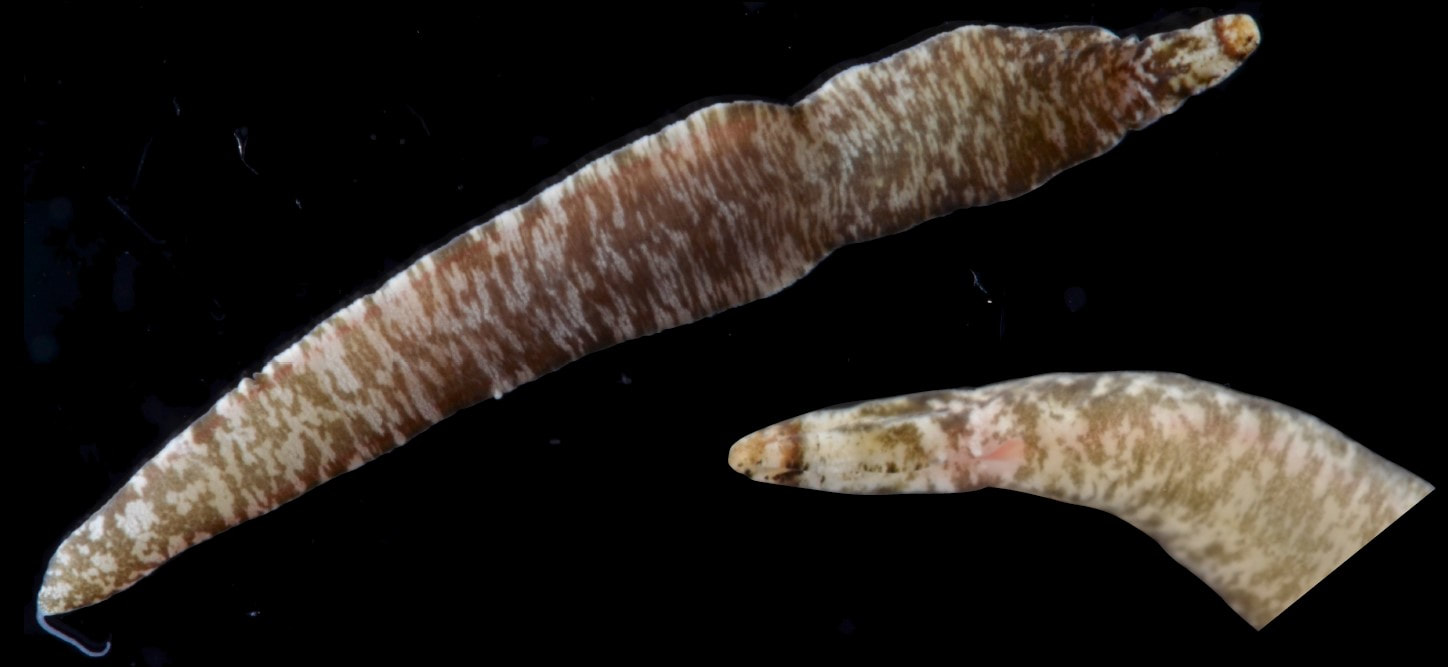
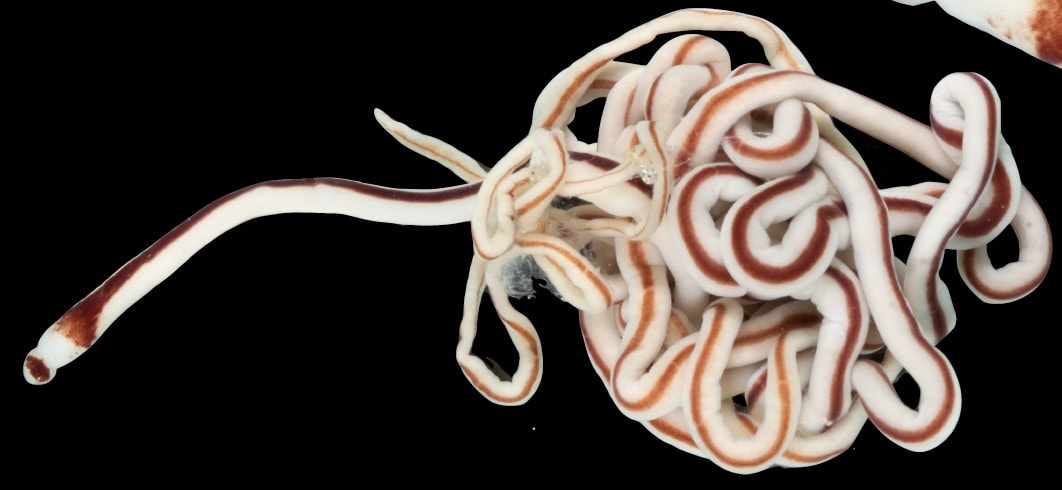
This is the crux of the project: by comparing genetic barcoding sequences to images which were taken in the field, we can look for ribbon worm species that are visually similar, but don't interact as one interbreeding species. Nemerteans desperately need this descriptive work because they have few notable external features except for color, which fades in preserved specimens. Also, these smooth, sliding, predatory worms with proboscides which pop out of their body cavities simply haven't received much attention. One of the best (and only) resources I've found so far about nemerteans in the Red Sea is from 1831, out-of-date in its descriptions and taxonomy, and written entirely in Latin. I never thought those four years of Latin classes could be so useful, so special thanks to Drs. Dombrowski, Beckelhymer, Martelli, and Goldberg! Next week, I'm looking forward to even more jigsaw puzzles, starting up the OIMBand -- I hear we have two violins, a ukulele, a guitar, and a banjo, so it's going to be awesome -- more lifer birds (I've seen Golden-Crowned Kinglets and Pigeon Guillemots for the first time), and learning more about the fascinating biology of nemerteans! |
|
Proudly powered by Weebly

















































































 RSS Feed
RSS Feed