|
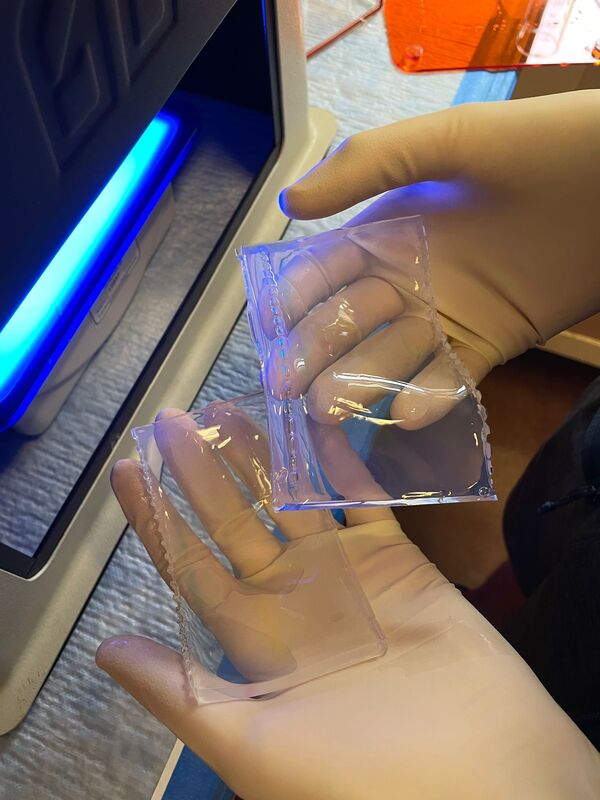
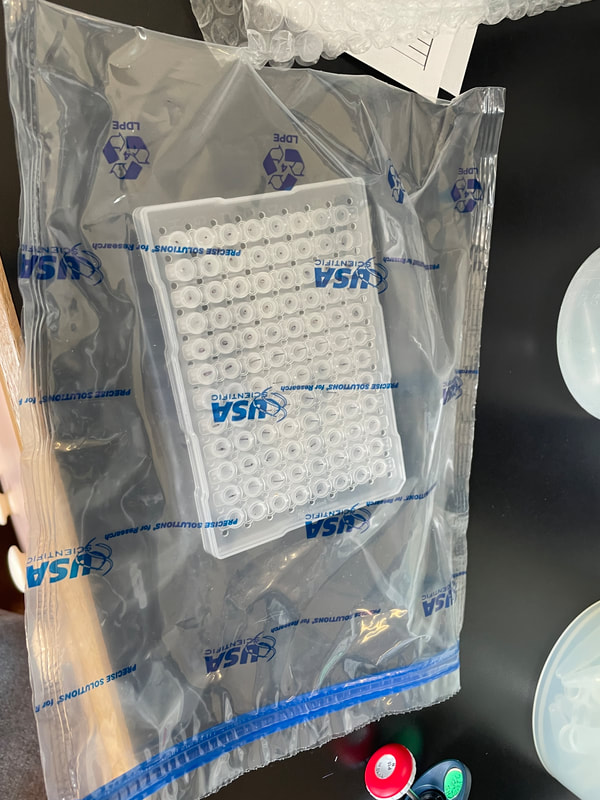
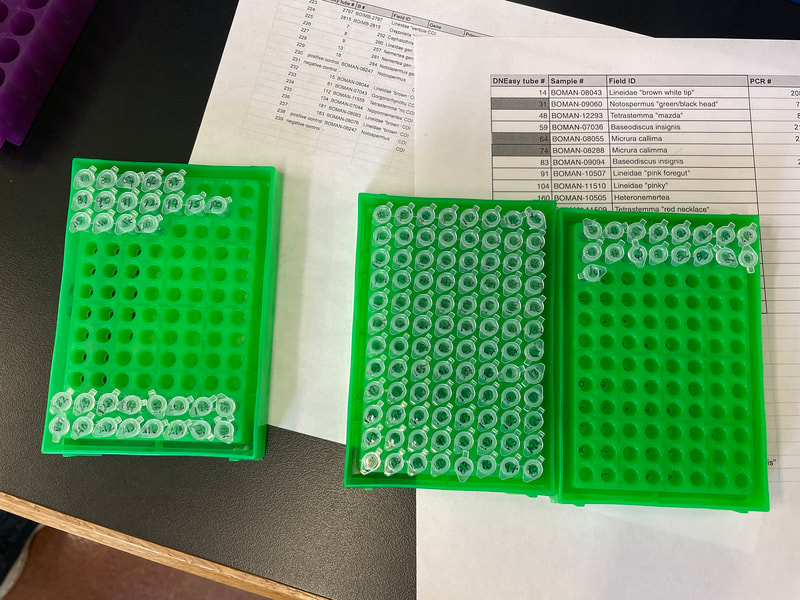
I can’t believe it is already week 5! The week started out great and slowly got more… well you will see when you read the rest of the blog. I didn’t get a chance to mention it in the last blog, but we saw the movie Barbie last Friday. It was highly anticipated among the interns since the beginning of the internship and it did not disappoint. We started week 5 with a boat trip on the OIMB’s new 48’ boat — R/V Megalopa! The weather was foggy, but the wind was only 10-15 knots, and the waves were relatively small. Check out the gallery to see what organisms we dredged up from 100-300 ft of water! Between looking down at the sea tables and the stench coming from the dredge, I got a bit sea sick. We also witnessed breaches in the water. One breach was from a seal, but we couldn't identify the second animal with certainty. We speculated that it might be another seal or its prey. At first, we considered porpoises, but their breaches were not consistent. This week in the lab was rough to say the least. It finally hit me how many times our PCR failed. Especially on Tuesday, none of the PCRs we ran worked. We have tried everything. From changing the primer combinations, decreasing the annealing temperature, diluting the samples, and doing a combination, none of the samples amplified. Some of the primer combinations, like LF/jgHCO, were new to us, and we only used them because we were running out of solutions and desperate for any progress. We've dubbed the 20 samples as our "problem children". We suspect that the problem might be our DNA concentration. However, theoretically, PCR doesn't require a large amount of DNA; it should work with just one strand. At this point, we are starting to feel like we're beating a dead horse if we continue to troubleshoot because it would lead us into an endless spiral of trying every possible combination of primers, temperatures, and dilutions. It would be a waste of time and resources to test out numerous combinations when the chances of success are low. The last solution to try is to re-extract if there is tissue left. It feels like we are treading water without making any progress and seem to be stuck in the exact same spot. We have made our way into the 600 zone for PCR tube numbers, 677 to be exact. This entire week Ethan and I stayed late in the lab hoping that our PCR would work. Our fruitful efforts were unfortunately not enough. There is more. On Thursday, we encountered a setback when the sequencing company, Sequetech, abruptly closed its operations after 30 years. As a result, we had to search for a new company to handle the sequencing of our samples. My mentor found a core facility at the Oregon State University's core lab, which seemed like a great option. However, there's a catch (as expected, it wouldn't be that easy). The DNA must fall within a certain concentration range and we need to pre-mix 1.2 ul of primer into the wells, totaling 12 ul. Unfortunately, it didn't end there. We couldn't determine the DNA concentration using our lab’s fluorometer (called Qubit), so we had to go with the second method, which is using a gel to estimate concentration based on the brightness of the amplified bands. We selected a range of samples to run on a gel and visually assess their brightness. However, since this process relies on subjective judgments, it introduces the potential for errors. If we fail to find another company to handle the sequencing, we'll be left with the task of reviewing all the gel images of the 96 samples we plan to send in order to estimate their concentrations. From there, we'll have to calculate how much we need to dilute each sample, adding further complexity to the process. To top it all off, I had high hopes that changing my laptop battery would work. To my disappointment, my laptop is still dead and useless. I think this is the trend this week: nothing is working :( We also had presentations this week. It went okay and it was fun to hear about the other intern’s progress. I was super nervous and I am glad that it is over. As a reward, all the interns went hiking after dinner! We went to the South Slough Trails and hiked to a saltmarsh. There were so many ripe blackberries, thimble, and salal berries. I really hope next week is better and we can send off the plates for sequencing.
0 Comments
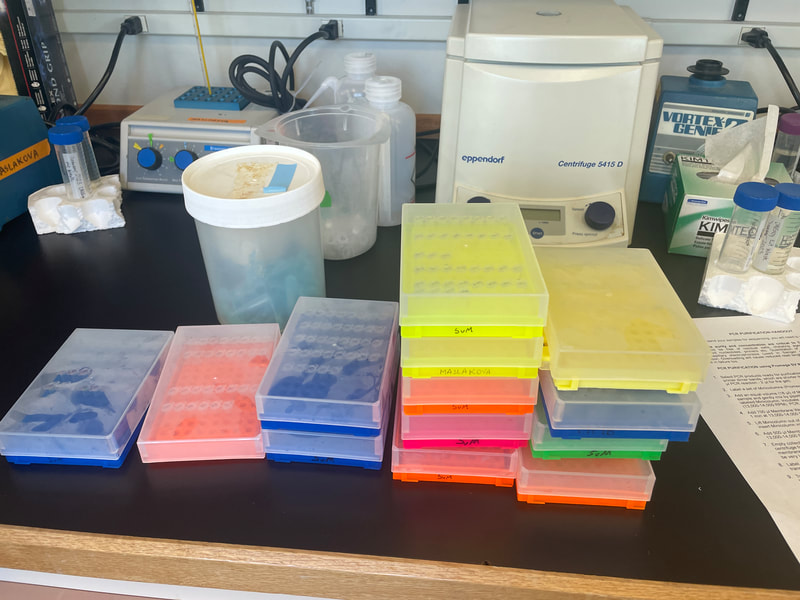
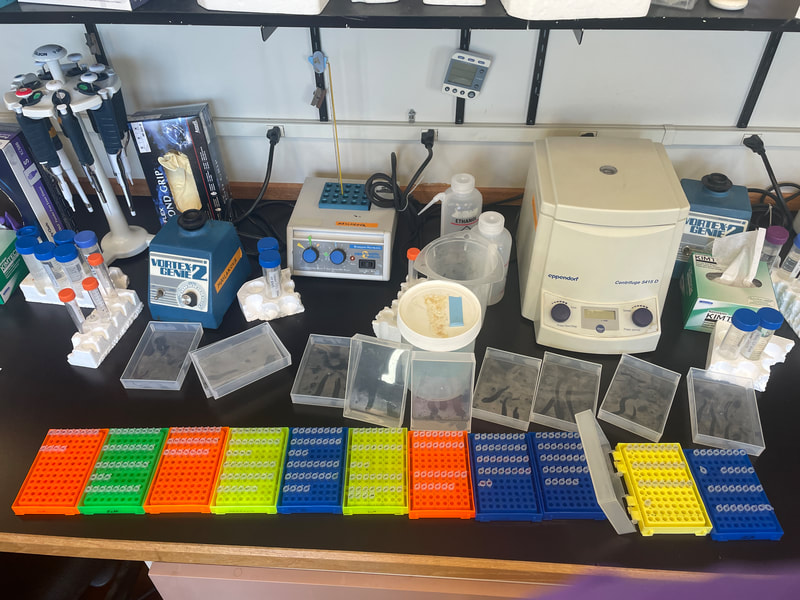
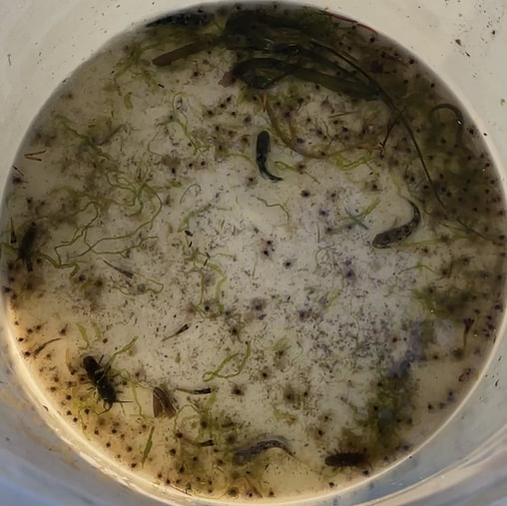
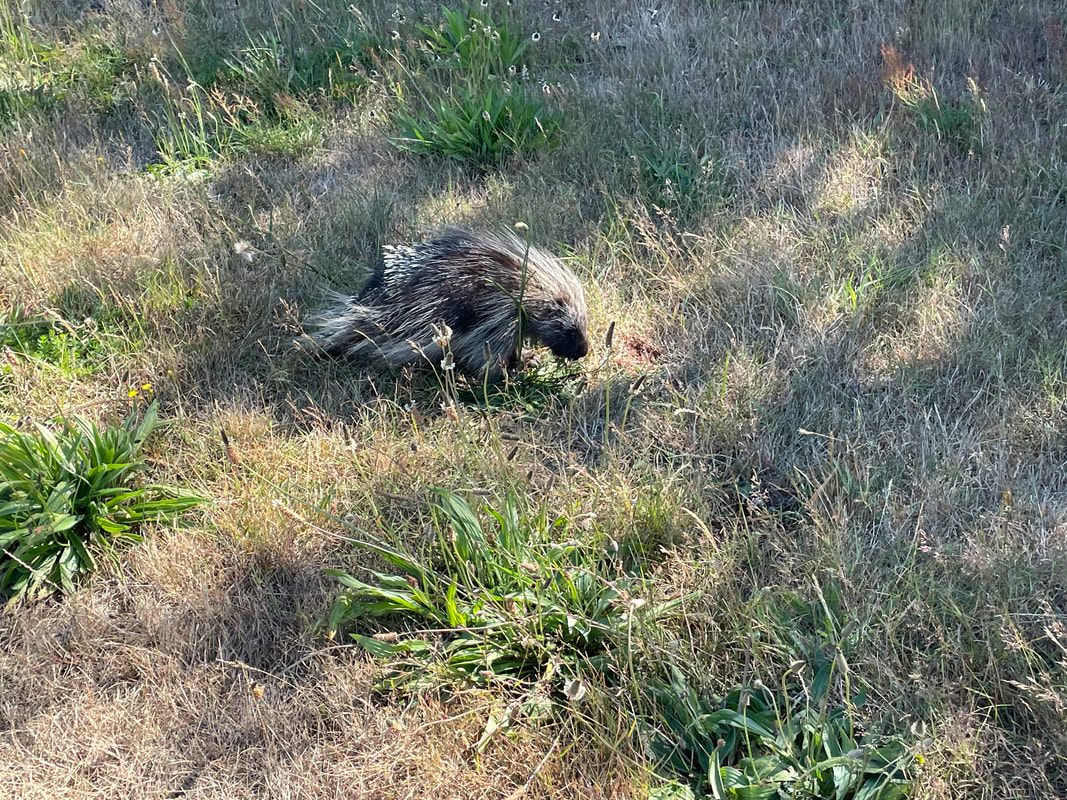
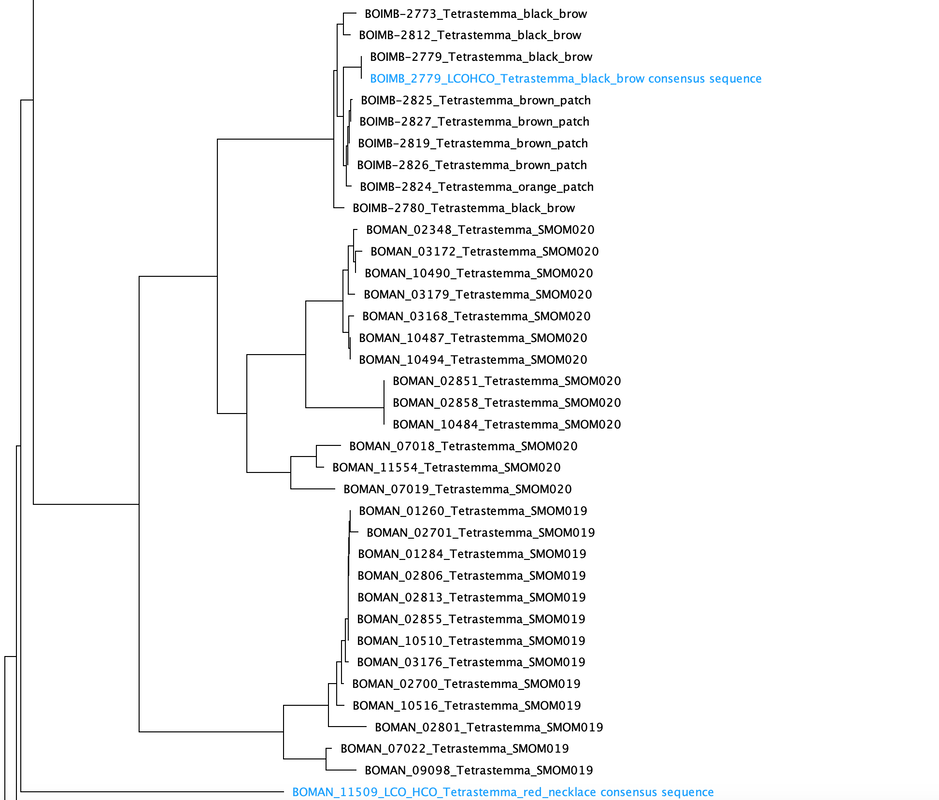
This week, the interns started off the week with a treasure hunt! We visited antique, vintage, and thrift stores at Coos Bay Boardwalk, where everyone found items that suited their style. As my mom would say, it was a productive trip! Personally, I love tiny things, particularly china teacups and trinket boxes, so I am definitely looking forward to returning. We also went back to the 7 Devils Pub and Brewery to see a band called Three for Silver. I personally liked this band better than the last band. Oregon State University interns came for a tour of our campus and the Charleston Marine Life Center (CMLC). We gave them a tour of our lab and tagged along the CMLC trip. On Monday, I joined the 20-plus-year-long larval crab project led by the OIMB Professor Emeritus Dr. Alan Shanks. We went to the docks to collect what was in the traps, recording the numbers of the Dungeness crab megalopa larvae and any other bycatch we came across. The catch turned out to be quite diverse, including isopods, rockfish, and shrimp! It was my first time at the docks and I was excited to see so many jellyfish there. Another new experience was plankton towing! Chloe allowed me and Ethan to join her for a plankton tow at the docks. We also went digging in the mud flats, where we discovered various organisms, such as clams, tube worms, phoronid worms, polychaetes, annelids (shimmy worms), and an ascidian. In the lab this week, we received more samples from Oman, courtesy of my mentor's colleague, Dr. Gustav Paulay. Just like the previous weeks, we extracted DNA, performed PCR, troubleshooted, ran the PCR samples on a gel, and purified the samples. We've already processed PCR tube number 400, and we're getting close to 500 :0! I wonder if we'll reach 1000 PCR tubes by the end of the project. Last week, we sent our samples for sequencing, and this week we got the results back. We found 7 new operational taxonomic units (OTUs — putative species) from Oman and Red Sea! Among them, there was a new species from the genus Gorgonorhynchus , 2 new species of Baseodiscus, 1 species from the family Lineidae, 1 Reptant polystiliferan, 1 species of Nipponnemertes, and 1 species of Tetrastemma. We believe that we'll discover many more Tetrastemma species, and overall, we expect to find numerous new OTUs. Unfortunately, my laptop broke this week, which couldn't have come at a worse time, as my proposal was due on Wednesday and I had a lot of edits to make. I made efforts to work on it in the lab early and after dinner. Additionally, I've been compiling all the species found in the Indian Ocean, which requires a lot of computer work. I have never done something like this before so it makes it a little inconvenient to have to reopen my sources every time I want to add to the list. Hopefully, after replacing the battery, my laptop should be up and running again! On Thursday, we were informed that Wallace the friendly porcupine that had been hanging around the OIMB campus had been removed by the Oregon Department of Fish and Wildlife (ODFW) to be relocated to the nearby hills. Sadly there won't be any more weekly Wallace photos to bless your feed. Compared to other weeks, this one feels less eventful, but time is flying by faster and faster! The highlight of the week was discovering numerous new OTUs, and I'm excited to send off another plate of samples for sequencing (hopefully next week). We started the week three off with camping at Camp Arago for 2 nights. On Saturday morning we went tidepooling and found a bunch of cool animals. My favorite was Antiopella fusca, a white-and-orange-tipped nudibranch. I was so surprised because I thought they were typically found in eel grass or on substrate, but this particular nudibranch was floating on the surface. Additionally, I saw a lot of bright orange-red nemerteans, Tubulanus ruber, and collected two to bring back to the lab. I saw lots of starfish (Pisaster ochraceus), purple urchins (Strongylocentrotus purpuratus), and a green anemone. We also went hiking and spotted some seals, a salamander, and several crabs. I grilled hot dogs for lunch and prepared hamburgers for dinner. The plan was to wrap up the camping trip by watching the sunset from the beach, but we got carried away digging holes in the sand in search of critters, paying little to no attention to the sunset. A few lessons I learned from camping: always remember to bring a pillow, make sure to have padding for your bed, and have enough layers or blankets as it can get really cold. Our sequences arrived this week! I learned how to process the sequences using Geneious Prime. First, the primers get trimmed, and low quality bases are removed from the ends. If there are two strands (one for each primer), they are aligned to make a double-stranded DNA. Then, a consensus sequence is created from the aligned strands and compared to sequences publicly available through GenBank, a database maintained by the National Center for Biotechnology Information, to find the best match. This step helps us confirm whether we indeed got a sequence from the phylum we expected (e.g. a nemertean, and not an annelid or crustacean, common prey items of nemerteans). A tree was constructed based on the relationship between the neighboring samples and provides us with a visual representation of the identified Operational Taxonomic Units (OTUs) and their relationships to one another. This also helps us determine which samples are cryptic species (there are a lot!). I was really excited to continue troubleshooting PCR, as we were able to send off another plate this week! I can’t wait for the results to come back so that we can add more individuals to the tree. Some of the previously sequenced samples showed amplification of a prey item, requiring us to try different primers. Our hope is that these new primers will better match nemertean sequences. Additionally, we encountered the same issue this week as we did last week with the presence of double bands. We have two potential solutions: first, purifying the PCR solutions in the hopes that it won't affect the sequencing process; or second, increasing the annealing temperature to eliminate the fainter second band. I started working on my project proposal this week, and it is progressing okay. However, I'm uncertain about the level of detail and development required due to the page limit. Fortunately, since we began searching for citations in the first week, it reduced the workload for this week. My mentor has also provided us with awesome resources and an outline to follow, which has been super helpful. She is the best! While searching for citations, I got to read more about the biotoxins found in nemerteans. Some of these toxins include tetrodotoxin and neurotoxic peptides. Interestingly, other organisms such as the blue-ringed octopus and pufferfish also possess tetrodotoxin. It is believed that this toxin is produced by endosymbiotic bacteria. Tetrodotoxin is 1,200 times more potent than cyanide, functioning by blocking sodium channels and interfering with the transmission of signals from nerves to muscles. So cool! I hope to learn more about it. Next week, we expect to receive more samples, and the process will repeat: DNA extractions, PCR, PCR troubleshooting, gel imaging, and sequencing. To start off the second week, the REU interns went kayaking at Hall Lake and climbed up sand dunes. Fortunately, the weather had warmed up just in time to go swimming! This week in the lab, I continued performing PCR. However, I noticed that my positive control was faint, making it difficult to observe. The positive control is a previously successful sample used to assess the viability of the master mix. This issue arose on two separate occasions, and I'm uncertain about its cause. To ensure thorough mixing of the DNA, which is not homogenous, I made sure to flick the positive control. Additionally, I fully incorporated the sample into the master mix and centrifuged it using the tabletop centrifuge. After discussing the matter with Ethan, we decided to switch our positive control to a more recent sample taken from the first PCR we conducted. We hoped that the positive control bands would be more pronounced, and indeed, they were brighter. Aside from the finicky positive control, I conducted PCR troubleshooting due to samples failing the initial PCR run. There are a couple of methods available for troubleshooting. The first approach involves diluting the DNA by using a ratio of 1:10 of DNA to nuclease-free water, which helps decrease the presence of PCR inhibitors. We found that a good amount of the samples worked after dilution. In some cases, an additional dilution of 1:100 may be necessary. Some of the samples yielded positive results using this method. The second troubleshooting method includes changing the primers. Currently, we are using Folmer primers, which serve as universal primers for a standard barcoding region of the Cytochrome Oxidase I gene for many metazoans. However, there are instances where the primers may bind to the DNA of a prey item in the gut instead of the predator (the specimen of interest to us), so in case of nemerteans one sometimes gets an annelid or a crustacean sequence instead. For the samples that were unsuccessful after the dilution, we tested out different primer combinations. The LF/ DR primer and LF/ HCO combination failed for most samples we were troubleshooting, so we are trying LCO/ DR and a set of degenerate primers. Since we had enough purified samples this week, we loaded them into a plate and shipped them off for sequencing. I am super excited to get the results back! My literature search has been yielding more progress since I expanded the scope of my search to the entire Indian Ocean, rather than just Oman. Unlike other species, nemerteans are not a hot topic in marine biology; however, they are in desperate need of attention. Ribbon worms, as top predators, play a crucial role in ecosystems by feeding on other organisms, like crustaceans and annelids. At most, the literature might contain descriptions of a couple dozen species, but many still await discovery, description, and classification. I am always amazed by how much science has developed. It used to be that histological sections were the best way to assess nemertean morphology and classify species, because the color of the live nemerteans would fade away after preservation. Now, DNA sequence data is routinely used to distinguish between species. I have always wondered how scientists developed procedures such as DNA extraction and PCR and how they determined the necessary chemicals for the reaction. This week, we went fossil hunting. The interns were hiking to a beach when we ran into a man carrying a fossil. At the beach, we were searching for fossils and only managed to find a small one. When we left the beach, the man had left the fossil on the trail with a note saying "Free." It felt as if it was meant to be, so we traded and left the small fossil for someone else to discover and keep. Additionally, at the same beach, we dug a channel to connect the freshwater with the ocean. However, the freshwater kept drying up, so we had to dig deeper as we approached the water. In the end, it was successful! On Wednesday, I attended my second seminar where two PhD candidates talked about their experiences on a research cruise. They even brought jellyfish for us to hold and pet, definitely one of the highlights of my week! Next week in lab we are continuing to do PCR troubleshooting, DNA extractions, and hopefully get to data analysis! I am excited to see the progress of the project. As a brief introduction, hello! My name is Audrey and I am majoring in Marine Science and minoring in Biology at Boston University. Ever since I was little, I thought cuttlefish and octopus were fascinating and have been interested in Marine Biology ever since. I hope to explore regeneration and stem cell research in marine organisms in the future. At Boston University I had the opportunity to work in a population genomics lab conducting research on coral to determine if the symbiont composition is impacted by environmental variation and if it affects the thermal tolerance of coral. I also worked in the wet lab feeding coral and conducting water quality tests. This summer, I will be conducting Polymerase Chain Reactions (PCR) on the Cytochrome Oxidase subunit 1 (CO1) gene under the mentorship of Dr. Svetlana Maslakova. Through the use of genetics, we aim to improve the knowledge of nemertean species diversity in Oman and the Red Sea, and to distinguish between cryptic species - organisms that share similar appearances but belong to different species. This week, Ethan and I completed 68 DNA extractions, consisting of 48 samples from Oman and 20 samples from the Red Sea. The extraction process involved lysing the tissues, binding the DNA to spin columns, and eluting the solutions to obtain liquid-form DNA. Unlike the other REU interns, our samples were already collected by our mentor. Next, we performed PCR to amplify a specific gene, the Cytochrome Oxidase subunit 1 which is a mitochondrial gene. CO1 is a highly conserved gene with specific regions that vary enough among species, allowing us to differentiate taxa. To assess the quality of the samples, we conducted gel electrophoresis and examined the resulting gel images. Additionally, negative and positive controls were used to detect contamination and evaluate the master mix.The successful PCR samples were purified. Next week we are aiming to send off a plate to get the samples sequenced. Aside from lab work, we were given readings and videos to enhance our understanding of the classification of nemerteans and their distinguishing features. There are three classes in the phylum Nemertea: Hoplonemertea characterized by stylets in their proboscis, Palaeonemertea, and Pilidiophora which is named after the pilidium larvae. One scientific article I read written by my mentor discussed how a nemertean species Micrura alaskensis is actually a complex of five different but cryptic species. I am currently in the process of finding literature, and I must admit that it is proving to be quite challenging. The topic I am exploring is largely unexplored, and narrowing it down to a specific location adds to the difficulty. During my freetime this week, I went to Davey Jones Locker, the pottery store, the OIMB beach, and the Charleston Marine Life Center. If I had more time, I would have volunteered in their aquarium. The campus has a lot of animals roaming around, so hiking is super fun. The REU interns and the summer students have a synchronized schedule because the meals are served for only 30 minutes. Breakfast is served at 7:00 am, which is something that I still have to get used to, but it makes the days seem a lot longer and more work can be done. All the interns check in with each other’s progress and it is so fun to hear what the others are doing. I wish there was more time so that I could spend nine weeks on each project. I would also like to go to the mudflats and experience collecting samples. I am excited to go on more hikes and check out the docks to see if I can find harbor seals. I am also looking forward to starting data analysis. |
AuthorHi everyone! I am Audrey and a rising junior at Boston University majoring in Marine Science and minoring in Biology. I am super excited to be participating in Dr. Svetlana Maskalova’s nemertean research this summer. Over the nine weeks, I hope to expand my experience with PCR. In my free time I like to go hiking, sewing, making jewelry, and baking. ArchivesCategories |
Proudly powered by Weebly














 RSS Feed
RSS Feed