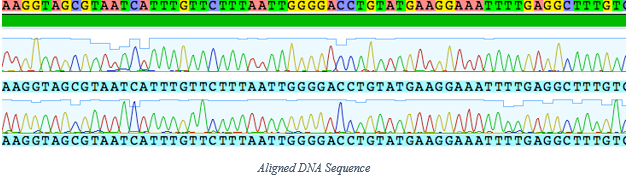
Week 8 To finish up my discussion regarding the processes involved in my research, this week I will cover sequencing the DNA. Once the DNA has been purified and quantified (week 7 blog post), we send our samples to a company called Sequetech located in Mountain View, CA in order to be sequenced. Once these sequences are ready, we need to analyze them and clean them up a bit. To do this, we use Geneious®, a sequencing analysis software. We first “trim off” the primer section of the DNA, then align the two complimentary strands to look for any discrepancies. Once the two complimentary strands are cleaned up and properly aligned, we can search for any matches to that particular sequence using GenBank®, a collection of publicly available DNA sequences. This data allows us to build phylogenetic trees and observe genetic relationships between species that we are studying. From here we are able to see if we have a species match, an undescribed species, or possibly even cryptic species. The past few blog entries have summed up the majority of my specific research, but the data that has been collected will be further analyzed and used as part of a much larger body of research long after I am finished with this program. I am excited to have the ability to follow the progress of this research into the future. 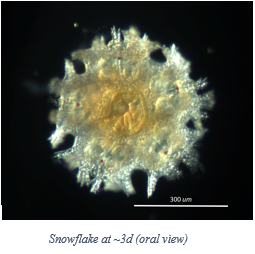 In week 2 I had mentioned that we had started some cultures of various organisms as a kind of side project, and this last week we have been able to see the results as our larvae have begun metamorphosing into their juvenile forms. The one that I am most proud of though, is our Patiria miniata larvae, as this is one of the more difficult organisms to induce to settle in a lab setting. When this program first began, I was informed that this had not yet been done in our lab, which of course I took as a personal challenge. I am excited to say that we have been wildly successful in getting our Patiria to settle, and I would like to introduce to you Snowflake: the first of his name, king of the larval culture, ruler of the Pyrex dish, and possessor of many tube feet.
0 Comments
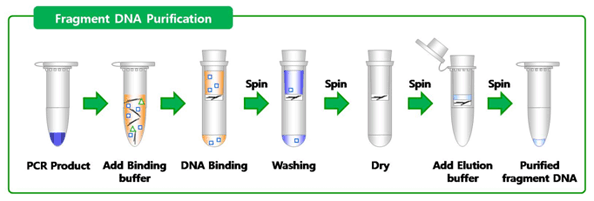
Week 7 This week I will cover the basics of DNA purification and quantification. Once we have amplified the target DNA using PCR, we need to purify the product (by removing primers, unincorporated nucleotides, polymerase, and salts) and quantify it in order to get a general idea of how much DNA is in the sample; if there is not enough DNA or too much DNA sequencing may fail. I used a commercial kit (Promega Wizard® SV Gel and PCR Clean-Up System) to purify our samples. First, a membrane binding solution is mixed with the PCR product and the mixture is pipetted into a spin column containing a silica membrane placed inside a collection tube; this assembly is then centrifuged in order to bind the DNA to the membrane (while everything else flows through into the collection tube). An ethanol-based wash solution is then added and pulled through the membrane by centrifuging in order to clean the filter. Once the sample is bound to the membrane and washed of contaminants, the spin column is placed into a fresh collection tube, and DNA is eluted by adding nuclease-free H20 to the filter, and centrifuging the assembly. This purified sample is then quantified using gel electrophoresis that I mentioned previously (see my previous blog post week-6.html). With this round of gel electrophoresis however, the sample is compared to a standard called Low Mass Ladder (which contains a mixture of fragments of DNA of known size and quantity) in order to estimate the quantity of DNA contained within the sample. Once the sample is purified and quantified, it is ready to send off for sequencing. The process of sequencing will be the focus of next week’s blog entry. (Photo courtesy of Bioneer Pacific) With only two weeks left in the program, we have officially run out of time for our usual weekly extracurricular activities. Focus has now shifted to compiling the data and preparing to present it to various audiences. We are all ready to see our hard work turn into something tangible, and the excitement around campus is palpable (punctuated with healthy doses of anxiety).
|
AuthorMy name is Becky, and I have been local to the Coos Bay and Charleston area for two years with my boyfriend David, and my dog Mojo. I moved to this area to complete my associate's degree at Southwestern Oregon Community College in anticipation of transferring to a four-year university. I now attend online at Oregon State University and I am planning a move to Bend, OR in August to continue at the OSU Cascades campus in order to finish my bachelor's degree in either natural resources or biology. Archives |

 RSS Feed
RSS Feed