 Slowly but surely, we are finishing up the PCR portion of the project. There are a few nemertean DNA samples that we are still struggling to amplify, but we have also started to analyze the data we have received from the company that sequences, or reads, our DNA barcode segments. When the DNA segment is sequenced, the data that we receive is in the form of two chromatograms per sample. We get two results per sample because DNA is double stranded, and the reaction that we use to sequence DNA, Sanger sequencing, only reads one strand at a time. We take these two complimentary strands and line them up using software called Geneious to ensure that they match, and no mistakes have been made. Once we have lined up the two strands in a “consensus sequence”, we check the sequence against Genbank. Genbank is a public, online database of DNA barcoding segments and the organisms that they belong to used by scientists all over the world. If another scientist has found the same species of worm as us and entered it into Genbank, we will find that our DNA sequence matches and know the identity of our sample. Unfortunately, most of our samples represent species that have not even been described, so we get very few species-level Genbank matches. We then turn to the database of Caribbean nemertean DNA sequences that the Maslakova lab has been compiling for the past few years. We compare our sequences to those that have already been categorized by Dr. Maslakova and her colleagues and are able to identify many more worms. The best part is when our worms do not match anything from Genbank or the unpublished Maslakova lab database. When this happens, we have found a new species of nemertean! So far, of the approximately 75 samples that I have sequenced for the gene CO1, I have found 30 different OTUs (operational taxonomic units, or what we call a species before it is officially described). Of those thirty OTUs, NINE represent new species that have never before been sequenced! I can’t wait to see what I find in my last 25 samples. Last weekend we visited Oregon State University’s Hatfield Marine Science Center in Newport, Oregon. All of us enjoyed watching a giant pacific octopus feeding at their visitor’s center as well as seeing their classroom and lab facilities. They have a particularly cool room dedicated to the microalgae that they feed their oysters with giant tanks of different algae species that were all different shades of green and brown. While in Newport, we also visited the Oregon Coast Aquarium where we all geeked out about the cool marine animals.
0 Comments
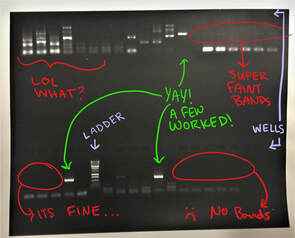 We are past the halfway mark, and everyone’s projects are in full swing! Adrian and I are spending our time in the lab troubleshooting pesky DNA samples. This is the less glamorous side of DNA barcoding. Sometimes, PCR reactions do not work correctly, meaning they do not amplify the correct section of DNA, on the first try. This requires changing the variables of the reaction, one at a time, little by little, until you finally get a enough of the same segment of DNA that is the right length. Why is the length of the DNA segment important? The length of the DNA segment is how we tell whether we have amplified the correct section of DNA, in my case the CO1 gene. We use a technique called gel electrophoresis. This method begins with placing the PCR-amplified DNA in the well of a gel. Next, an electrical field is introduced. Because DNA is negatively charged, it is moved through small pores in the gel by the electrical field. The smallest DNA segments move the fastest through the gel’s pores while longer DNA segments take more time to be pushed through the gel. When we take a picture of the gel after some time has passed, we know that the segments of DNA that have traveled the farthest, are the smallest. We use a “ladder” that is full of DNA segments of known sizes to compare our samples to and quantify their lengths. Using this technique, we can tell which PCR reactions worked (there is a band of DNA approximately 658 base pairs long – the size of the gene CO1) and which did not (the band of DNA indicates a segment too long or too short, there are multiple bands, or there is no band at all). We REALLY want there to be a single, bright band of the correct size, but if there is not, we keep trying until we achieve that result from our gel electrophoresis. We spent some time crabbing in the evenings this week. It turns out permits very inexpensive for Oregon residents (thank you Steven) and you can catch quite a few crabs, mussels, and other shellfish. It was fun to watch the crabs and throw the smaller ones back. Renee is teaching us all about crabs because her project requires catching a lot of them. She is doing a predation experiment where she observes which species of smaller crabs (Dungeness or invasive green crabs) that a larger species (red rock) prefers to eat, by putting them in containers and waiting until they eat each other.  Week four flew by! We have been going nonstop since last Friday when we began our camping trip at nearby Cape Arago State Park. That night was filled with s’mores, laughs, and songs around the campfire. A few of my fellow interns are quite talented on the guitar and ukulele! On Saturday we checked out North Cove. It was a great new site for tide pooling, and we learned a lot. It amazes me that every time I go out to the tide pools, I see so many new organisms. On Saturday, we were joined in our camping endeavors by University of Oregon SPUR students. SPUR students are interns on main campus this summer, and they came to see what OIMB is all about. We had a great time getting to know them and playing some ultimate frisbee, and on Sunday we once again went tide pooling (which we would do every day if we could – it never gets old!). Our big find of Sunday was an octopus. We loved all the festivities but also had research proposals on our minds. On Tuesday, we submitted research proposals detailing the what, how, and why of our projects. Having a clear plan in place to answer our research questions is great, but it was also very helpful to step back and look at the big picture of why our project is important to everyone – not just to those of us who love look at and learn about worms all day. In the lab, we continued PCR and sent a big batch of samples out to California for sequencing. We can’t wait to get the DNA sequences back and find out which species of worms we have been working on for the past couple weeks! Week four also meant that we had to say goodbye to the graduate student who spent the first three weeks of this summer teaching my lab partner Adrian all about nemerteans, PCR, DNA barcoding, and so much more. We are going to miss Christina!  Week three was a busy one. This week we set up PCR reactions for many of our nemertean samples. PCR stands for polymerase chain reaction. It is a process widely used in biological research that allows us to select just a small piece of an organism’s large genome and make tons of copies of it. This amplification of a small portion of DNA is essential to get enough copies of the DNA segment to have it sequenced so that we can read what the DNA says. The small DNA segment I am focusing on is the protein coding gene, cytochrome c oxidase subunit 1 (CO1). This is a gene that is found in the mitochondria of a cell rather than the nucleus. CO1 is a widely used gene for DNA barcoding because it mutates fairly quickly but has sections on either side of the gene that have stayed almost the same. Because the edges of the gene segment are the same in most animals – from worms to humans – we can use those “conserved” sequences to find the gene and amplify it with PCR. Using this particular gene has its advantages not only because it has conserved edges that allow us to find the gene, but also because other scientists studying all sorts of organisms all over the world also look at CO1. These scientists upload their CO1 sequences online so that we, and anyone else, can compare our DNA barcoding segments to theirs and see if we have the same species, a closely related organism, or something entirely different. It is an extremely powerful tool. After we amplify the DNA sequence that we want with PCR, we purify the DNA and send it off to be sequenced. Once we receive those sequences, we will compare them to the Maslakova lab database of Nemertean sequences from the Caribbean, and GenBank, which will help us verify the identity of the worms in our samples and establish whether we have any new species. We expect that we will find several new species (and have already found a few!) because the Nemerteans were collected from a location that has never before been sampled. This was a shortened week due to the Fourth of July, but it feels like we accomplished a lot in the days that we worked. On Friday, we got out of the lab for the morning and checked out a new tidepooling location… Middle Cove! We found more a few more Nemerteans to bring back to the lab as well as plenty of other fun invertebrates. The evenings have been filled with bonfires on the beach and hiking. Last weekend we went to Eugene to check out the University of Oregon main campus, and I think we all agreed we like it better out here at OIMB. This weekend we have a camping trip that everyone is getting excited about, so tune in next week to hear about that adventure. Nemerteans. I have learned so much about this phylum of predatory ribbon worms in the past two weeks. The group is defined by the synapomorphy, or trait that all species within the group have, of an eversible proboscis which they use to capture prey. The nemertean everts the proboscis, which is covered in toxic mucus secretions, and wraps it around its prey. The prey is slowly paralyzed and is then swallowed whole by the nemertean. In one group of ribbon worm , the proboscis even has one ore more small spikes or stylets, that inflict an injury to the prey that allows the toxic secretions to work more quickly. There are approximately 1,300 described species, but likely several times that number exist. This is because they are not well-studied and because many of these worms look a lot alike but are actually genetically distinct species . That is, they are “cryptic” species! That is where Dr. Maslakova’ s work comes in. As I mentioned last week, my lab partner, Adrian, and I are working on a long-term project in the Maslakova lab aimed at quantifying the diversity of the phylum Nemertea in the Caribbean. The first step in that process is to extract DNA from the worms’ tissue.
This week, we mashed up worms. We have around one hundred ribbon worms that were collected in Colon, Panama. Each specimen has to be ground up and combined with an enzyme to help break down the tissue. From there, we are able to extract the worm’s DNA. We finished these extractions for all of our Nemerteans this week and will amplify a small segment or barcoding region of each worm’s DNA next week. Comparing those barcoding regions will help us to determine how many different species exist in the area. In non-lab news this week, my fellow interns and I have been taking full advantage of our beautiful surroundings. On Sunday, we spent the afternoon at nearby Bastendorff beach where we worked on our frisbee skills, looked at crab molts, and went for a quick swim. Yes, a few of a were brave enough to run all the way into the extremely cold ocean only to quickly retreat (twice). Our nights this week have been spent finding new trails in the woods and watching the sunset on beautiful new beaches. The constant chatter about research, and a shared love of marine invertebrates keeps us happy, busy, and excited for the weeks to come. |
AuthorHi! I’m Megan Powers, a fourth year Biology and Environmental Sciences student at the University of Iowa. Throughout my summer at OIMB, I will be working with the Maslakova lab to assess Nemertean diversity in the Caribbean. Archives
August 2019
Categories |
Proudly powered by Weebly












 RSS Feed
RSS Feed