|
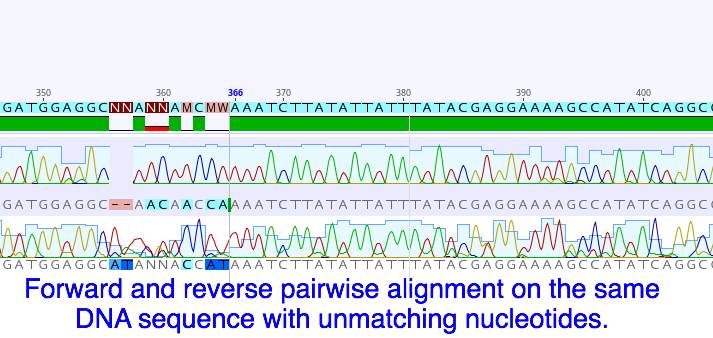
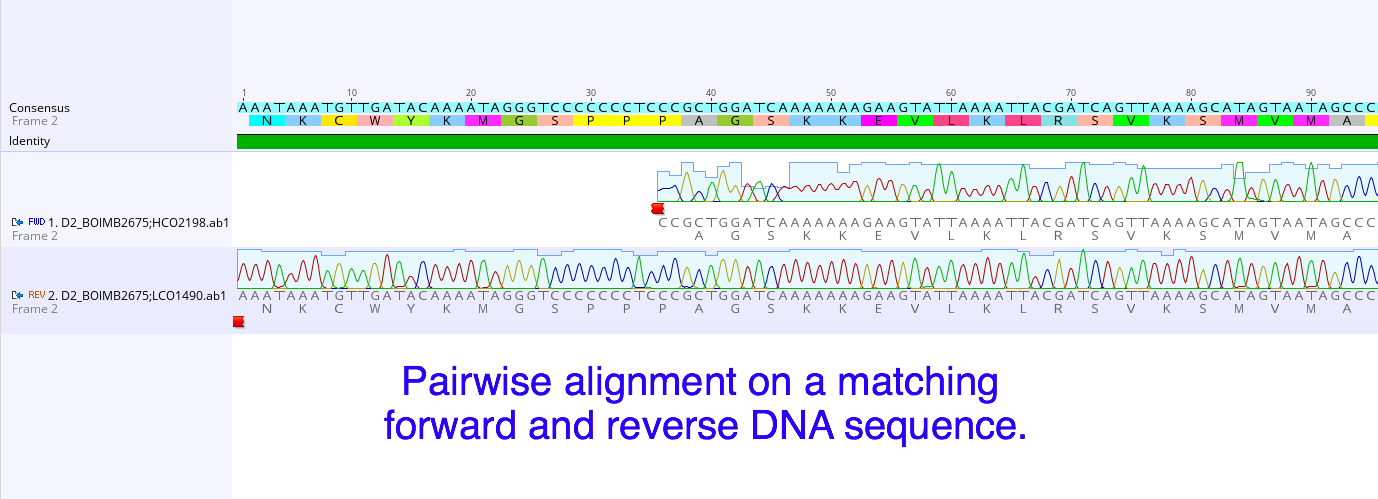
The DNA sequences came back for the first round of specimens. As I understand it, to create the sequences a process similar to the PCR is done but the DNA nucleotides are exchanged for chain-terminating fluorescent nucleotides. That means each one of the four kinds of nucleotides in DNA is tagged with a specific color and when they are incorporated into the growing chain, DNA synthesis stops. The fragments of DNA are then sorted by size on a gel (using electricity to drive the negatively charged DNA towards the positive end). The machine then reads the different colors and converts the colors into their corresponding nucleotides. Ta-da you now have the sequence of nucleotides for that DNA. This is obviously the super simplified version and of course a lot of things can go wrong along the way. For a deeper dive into this topic you can read "Sanger Sequencing Steps & Method" by Millipore Sigma. It is still possible the sequence will be unsuccessfully read. Even though we check the sample before it is sent out to make sure the DNA has been amplified, we have no way of knowing who's DNA was amplified. It is possible that the sample could get contaminated (don’t sneeze!) at some point during the extraction and amplification process. Another issue is the nucleotides in a sequence may not always come out clear enough to read with any certainty. This is why for each specimen we send a forward and reverse primer to initiate the sequencing reaction. This will give us two sets of the same sequence, one in each direction. Using a software program called Geneious we can overlap and compare the two sets for better accuracy. Working in Geneious has been really interesting. Once we trim up the two sequences, clean them up and merge them, we can then go on to the National Center for Biotechnology Information and upload the sequence to their program called the Basic Local Alignment Search Tool (BLAST). The BLAST program tracks uploaded sequences and their related species so we can use it to see if anyone has entered any information on known species that match our sequences. This part is exciting and really fun, most of the time it will also show you where the samples were collected from geographically. There is a lot more to this program and I’m looking forward to getting to know it over the next few weeks. Also this week we finished photographing and tagging all the worms we want to process from everything we’ve collected so far. We got all the DNA processed(extracted, amplified, and purified) and are hoping to ship out our second batch on Monday. In between that we have been able to look over and organize the BLAST information on the DNA sequences. The last round of field collecting should be this week also, which we’ll start processing as soon as we can. Hopefully we should have all the specimens processed and sequenced by the end of next week. Then we will be focusing on getting the information organized and displayed properly for the identification guide.
0 Comments
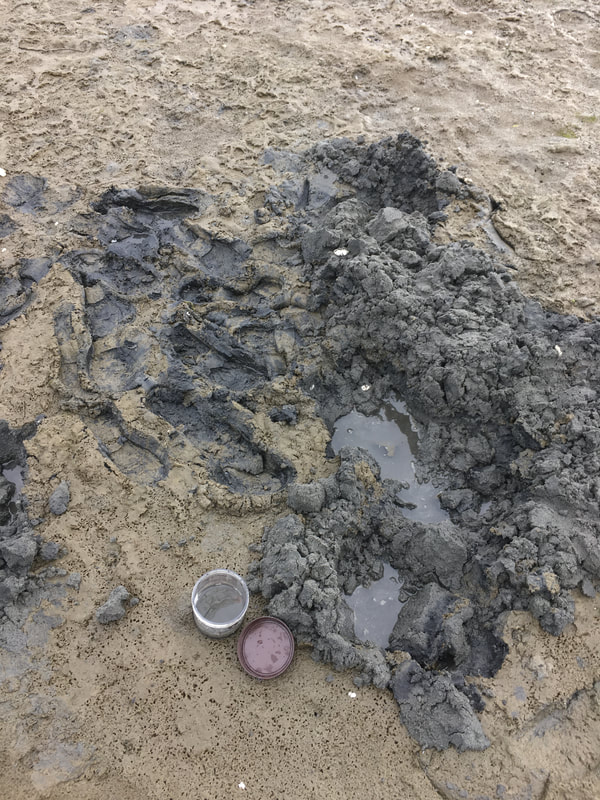
We have started collecting worms again! We spent Friday, Monday and Tuesday morning at various mudflats and sandy beaches. There are one or two species of nemertean worms that can be found on the surface but most of the worms found in this habitat are living in the sand and mud (which means a lot of digging!). Finding worms can be tricky and take a lot of shoveling. Hopefully if you are lucky enough to find one, they will be intact and not split in two by the shovel head. It can be really easy to break some of the worms when trying to get them out of the sand. So instead of pulling you have to carefully break the sand away away from each worm. It is really exciting when you do find a large one. They swim through sand like it is water.
One of my favorite genus of Nemerteans live in the mudflats, the Cerebratulus. They are larger then most of the other worms and can swim like an eel. They move their head like a snake but instead of eyes they have two longitudinal slits on either side of their heads called the cerebral organ furrows. These furrows are located next to two sensory organs and it is unclear which sense they might have and are using to understand their environment. Such as if these sensory organs can detect a sense of smell, taste or of movement in the water. They move their head in a peristalsis wave that gives you the impression that they are sampling the water in their environment. These slits are deeper and more developed then some of the other species of worms. They are overall very graceful and seem to move with intent. They change color in captivity and become limp and lethargic but will usually immediately revive and explore when their water is changed with fresh sea water.
Video of the Cerebratulus head waveVideo of Cerebratulus swimming in fresh filtered sea water.
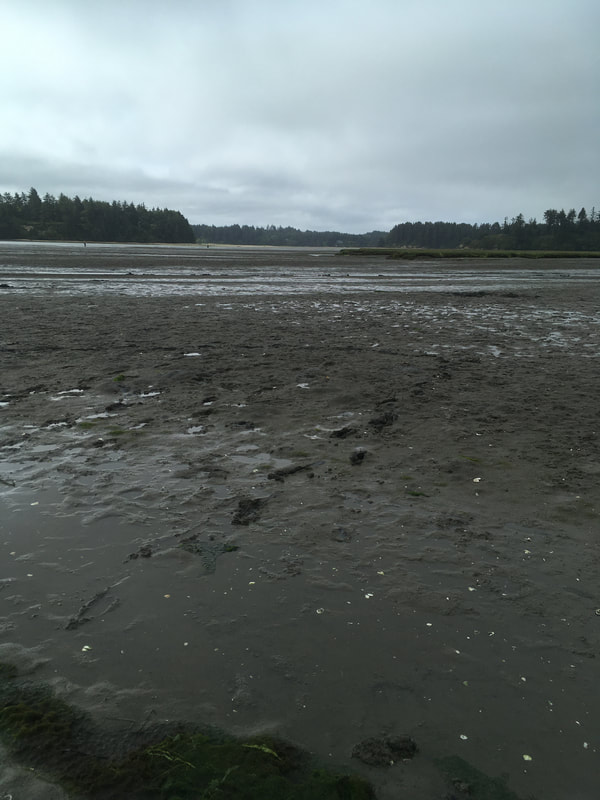
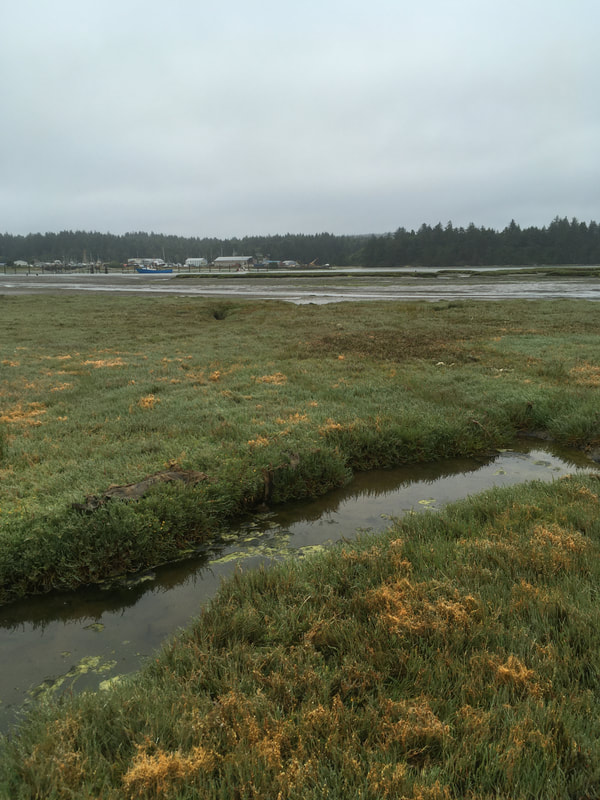
It is surprising to me the biodiversity found living in the sand and mudflats. Not only is their a variety of nemertean species but there are creatures of all shapes, sizes and colors. The biodiversity also changes quickly (within a matter of feet) from one location to the next. At first I thought all the mud and sand looked the same but after a few hours of digging your realize the subtle and no subtle changes in the sediment. Sometimes the sand and silt are dense, tightly packed and can even be slightly smelly. A shovel full might retain its shape or the sand might easily fall apart. I wonder if you sampled the sediment content vs the biodiversity if there would be any patterns.
I got to get up close and personal using the slr digital camera that attaches to a microscope and can take close ups of some of the smaller worms. I got a great view of the variety of features from the different nemertean species. I learned that adjusting flash and exposure on a moving object is a lot harder then it looks! This step, although critical to properly documenting each worm and its features, is definitely a bottleneck in the time it takes to process each worm. Yet, I do believe the end products prove that a picture is worth a thousand words!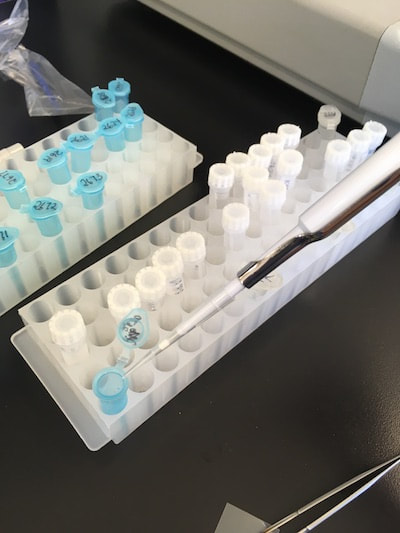  By the end of last week we had photographed and catalogued about 46 different worms. Which were then ready and waiting to have their DNA processed. So on Monday, Tuesday and Wednesday we were busy in the lab, pipette in hand, starting with the DNA extraction, then running the PCR, testing our samples to make sure the primers worked and lastly getting them ready in well plates to ship out this Thursday for sequencing. The protocols for each procedure creates a lot of “busy” work of just focusing and following one step after another. It has a nice, steady, and organized rhythm compared to the activity of field work and the patience of photography. The three tasks definitely hold an appealing balance between them all. Tomorrow we are going over how to interpret the sequences and use the software so that we will be better prepared for when we receive the sequences for our samples. I’m really looking forward to finding out how all that works! We have also met with various mentors at lunch who have all been truly amazing. Our professional development days have also been really useful and lately we have been working on making and giving presentations and how to begin making our scientific posters. The tips and advice have been extremely helpful. I will admit I was a bit nervous when I first realized we had to create a scientific poster on our project but I soon realized we are actually guided through the layout of how to put it all together. Now that my material is beginning to come in I am actually looking forward to putting it all together.
Wow, what an amazing last few weeks here at OIMB! I am surprised by what we have already accomplished and learned in such a short period of time. We have really hit the ground running to get started on the biodiversity project for nemerteans that are found on the Oregon coast. As I have now learned there are many parts to properly completing a modern biodiversity survey and creating an identification guide. Some of this process has been unchanged for hundreds of years, such as the observation and classification of different types of species based on their morphological features. Morphology is the study of the forms of things and has historically been the primary tool for identifying different species of organisms. The problem being that if you ask a group of ten people to observe something and then describe it, chances are there will be variations in each persons description. So as you might assume, this was never an exact science and there is lots of room for error and no way to “clean it up” with any amount of certainty, But now with DNA sequencing there is a means to more objectively identify and group different species. This not only helps to confirm specific features belonging to a specific species but can also alert us to cryptic species. Cryptic species are organisms that look alike. DNA barcoding can reveal that although these species can look very similar their DNA is, in fact, different enough for them to belong to different species, tells us they are genetically distinct . The opposite can also be true; some animals that appear to be very different, are in fact the same species but with varying color patterns. A Glimpse into Molecular Biology:)The first week here at OIMB we went through the steps for processing the DNA. Christina Ellison, a graduate student working with Dr. Svetlana Maslakova, our research mentor, guided us step by step through this fascinating process. The first step in this process was to extract and purify the DNA from the nemertean tissue samples. These first samples were collected prior to our arrival and placed in individual vials of 95% ethanol, which where then stored in a freezer at -80 ℃ to prevent tissue degradation. The first step was to extract the DNA from the chunks of tissue. This is accomplished using a miniature pestle to manually grind the tissue and by adding a lysis buffer solution. The lysis buffer helps to break open the microscopic cell and nuclear membranes, essentially freeing the DNA strands from inside their protective nucleus walls. A small amount of proteinase K is also added to the mixture to help digest the proteins and enzymes in the tissue. The Proteinase K keeps the naturally occurring enzymes, DNAse and RNAse, from doing their job of breaking down the DNA and RNA, thereby keeping the DNA intact for us to use for DNA-barcoding. The DNA then needs to be separated from the other cell debris. One way this is accomplished is by running the mixture through a special “DNeasy” column that has an artificial filter membrane. This process includes a variety of steps but essentially the DNA binds to the membrane in the vial so that the debris can be washed through it, leaving clean DNA strands attached to the membrane. The DNA is then released from the membrane into nuclease-free water, to be collected and used in the next step. Here comes the really fascinating part. In 2003, the Cytochrome C oxidase subunit I gene (commonly abbreviated as cox1 or COI) was proposed to be used as a universal gene region for DNA-barcoding of animals. This sequence changes fast enough over evolutionary time to differentiate between even closely species, but is also flanked by two regions of DNA that have not changed much across the animal kingdom. Therefore the markers on each end are very similar in most animals, and can be targeted to help amplify the highly variable region of DNA in between, regardless of wether you are sampling tissue from a rabbit or a nemertean worm. The key here is that you need to have the same region of DNA from a large number of organisms in order to be able to compare and contrast sequences across the animal kingdom. The DNA processing is of course not the first step in completing a biodiversity survey but by running through this procedure first it helped me to see the value of DNA barcoding in species identification, and cataloguing species diversity. It also made me aware of the importance of properly handling and labeling the tissue during field work and the importance of not contaminating the tissue samples with DNA from other sources. The Journey Begins!So, last week we officially began our first biodiversity survey. The first step of a survey is actually preparation and includes things like getting a collection permit, scheduling and preparing for the field work, planning for specimen storage and tracking. This step was already completed by our mentor, Prof. Svetlana Maslakova, who has been collecting and studying nemerteans in various parts of the world for the past 20-some years. The beginning of last week was spent getting familiar with the taxonomy and differentiating features of the various types of nemerteans. I admit the taxonomic names and pronunciations have been a bit of a challenge for me, but I am persistent and determined to keep trying!
The next step is field work, which for us meant a lot of early morning tide pooling and collecting nemerteans from various habitats. Enjoying the views and getting to know Southern Oregon’s diverse marine life was a fantastic way to spend the mornings and explore the coastline. It was important to collect as much information as possible for each individual worm, and it helped that our mentor has an established system of recording and cross referencing data from each trip. The tide pools provided a variety of habitats to sample from such as underneath rocks, in mussel beds, in the sand, on surface of rocks or among kelp holdfasts or surfgrass roots. Each location and habitat is recorded and given a reference number. Once back at the lab the worms are put into beakers and bowls and placed in the sea table as quickly as possible to stay cool. Then, they are grouped by morphology, identified to the closest known taxon, and labeled with location and habitat.. This made for a few busy but exciting days, and it was really fun getting to know the worms I have heard so much about. They were surprising and definitely have more personality then I would have ever thought. We have begun processing the worms this week. Every worm that is being processed gets assigned an individual number, and is documented with color photographs of its features. Some worms are large enough for a standard macrophotography (using an SLR camera with a macro lens), while others need to be photographed through a microscope. Learning how to work with the different camera equipment and microscopes has been a really cool bonus that I was not expecting and have thoroughly enjoyed. Once the specimens have been properly documented, they are preserved in two separate vials. Some of the tissue (usually the posterior end) is placed in 95% ethanol for DNA-barcoding and some (usually the anterior end, which is more feature rich in these worms) into 10% formalin to be preserved and later deposited into a natural history museum as a morphological voucher. The morphological voucher is a way to provide a reference specimen for other scholars to see the features first hand. |
Rebecca OrrHi, my name is Rebecca I’m from Northern California. My major is in biological sciences. I am so excited to be working in Dr. Svetlana Maslakova’s lab and learning the protocol and procedures for completing a modern biodiversity survey. Archives
August 2021
Categories |
Proudly powered by Weebly



 RSS Feed
RSS Feed